Team:Calgary/Project/OSCAR/Decarboxylation
From 2012.igem.org
| Line 36: | Line 36: | ||
BioBrick consisting of two primary genes. These include acyl-ACP | BioBrick consisting of two primary genes. These include acyl-ACP | ||
| - | reductase (AAR), which reduces fatty acids bound to ACP to fatty | + | reductase (<i>AAR</i>), which reduces fatty acids bound to ACP to fatty |
| - | aldehydes, and a second gene called aldehyde decarbonylase (ADC), | + | aldehydes, and a second gene called aldehyde decarbonylase (<i>ADC</i>), |
which subsequently cleaves the entire aldehyde group and results | which subsequently cleaves the entire aldehyde group and results | ||
| Line 54: | Line 54: | ||
| - | <p>This lead us to believe that the PetroBrick may have the | + | <p>This lead us to believe that the PetroBrick may have the potential to turn NAs in to hydrocarbons and be a perfect solution to remediating NAs! First though, we needed to show that the PetroBrick did in fact work as expected. We had some difficulty with the DNA from the registry and had to request the constructs directly from the Washington team. Once we had the Petrobrick, we needed to verify that the Petrobrick would work in our hands as it did for the 2011 Washington team. |
| - | + | ||
| - | potential to turn NAs in to hydrocarbons and | + | |
| - | + | ||
| - | be a perfect solution to remediating NAs! First | + | |
| - | + | ||
| - | though, we needed to show that the PetroBrick did in fact work as | + | |
| - | + | ||
| - | expected. We had some difficulty with the DNA from the registry | + | |
| - | + | ||
| - | and had to request the constructs directly from the Washington team. Once we had the Petrobrick, we needed to verify that the Petrobrick would work in our hands as it did for the 2011 Washington team. | + | |
Figures 2 and 3 demonstrate the function of the Petrobrick.</p> | Figures 2 and 3 demonstrate the function of the Petrobrick.</p> | ||
| Line 71: | Line 61: | ||
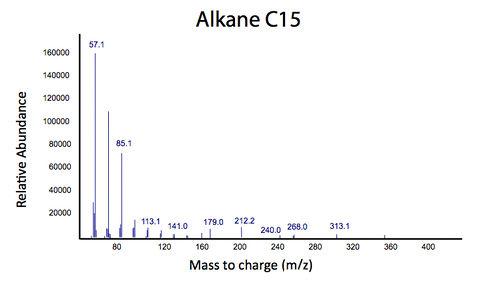
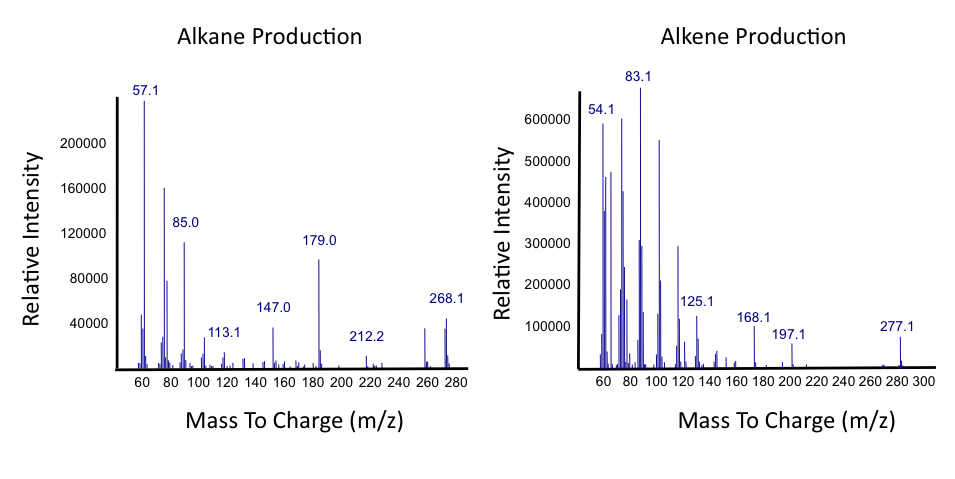
</html>[[File:Calgary2012_PetrobrickVerificationMS.jpg|center||thumb|Figure 3: Mass Spectra of the gas chromatograph peak at 12.25 min. The spectra suggests that the Petrobrick is selectively producing a C15 alkane. This is what was expected as determined by the Washington 2011 iGEM team.|500px]]<html> | </html>[[File:Calgary2012_PetrobrickVerificationMS.jpg|center||thumb|Figure 3: Mass Spectra of the gas chromatograph peak at 12.25 min. The spectra suggests that the Petrobrick is selectively producing a C15 alkane. This is what was expected as determined by the Washington 2011 iGEM team.|500px]]<html> | ||
| - | <p>With the Petrobrick shown to be able to successfully produce | + | <p>With the Petrobrick shown to be able to successfully produce alkanes, it was time to test it out on NAs, to see if |
| + | they could be selectively converted into alkanes! This experiment used commercially available NAs fractions including a large number of different complex NAs compounds. </p> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
<h2> Successful conversion of NA's into Hydrocarbons!</h2> | <h2> Successful conversion of NA's into Hydrocarbons!</h2> | ||
| Line 94: | Line 76: | ||
<h2><i>Nocardia</i> Carboxylic Acid Reductase (CAR)- Can we do better?</h2> | <h2><i>Nocardia</i> Carboxylic Acid Reductase (CAR)- Can we do better?</h2> | ||
| - | <p>Although we were successful using the Petrobrick to remove carboxyl groups from NAs, we wanted to improve on our results to see | + | <p>Although we were successful using the Petrobrick to remove carboxyl groups from NAs, we wanted to improve on our results to see if we could get a higher yield or possibly target other compounds. One of our original fears in using the PetroBrick to |
| - | + | decarboyxlate NAs was that the first enzyme AAR was reported to be highly specific for fatty acids bound to ACP. We had concerns about its compatibility with NAs and therefore sought another enzyme in the literature called carboxylic acid reductase (CAR) that was documented to perform a similar task as AAR, converting fatty acids to aldehydes, but with much lower specificity (He et al, 2004). This enzyme, from <i>N. iowensis</i> does not require covalent attachment to ACP so would | |
| - | if we could get a higher yield or possibly target other compounds. | + | likely be much broader in substrate specificity. It requires a second gene from <i>N. iowensis</i>, called Nocardia phosphopantetheinyl transferase (<i>NPT</i>) necessary to append a 4’- phosphopantetheine prosthetic group to CAR required for its full function (Venkitasubramanian et al, 2006).</p> |
| - | + | ||
| - | + | ||
| - | + | ||
| - | decarboyxlate NAs was that the first enzyme AAR was reported to be highly specific for fatty acids bound to ACP. We had concerns about its compatibility with NAs and therefore sought another enzyme in the literature called | + | |
| - | + | ||
| - | carboxylic acid reductase (CAR) that was documented to perform a | + | |
| - | + | ||
| - | similar task as AAR, converting fatty acids to aldehydes, but with | + | |
| - | + | ||
| - | much lower specificity (He et al, 2004). This enzyme, from <i>N. | + | |
| - | + | ||
| - | iowensis</i> does not require covalent attachment to ACP so would | + | |
| - | + | ||
| - | likely be much broader in substrate specificity. It requires a | + | |
| - | + | ||
| - | second gene from <i>N. iowensis</i>, called Nocardia | + | |
| - | + | ||
| - | phosphopantetheinyl transferase (<i>NPT</i>) necessary to append a 4’- | + | |
| - | + | ||
| - | phosphopantetheine prosthetic group to CAR required for its full | + | |
| - | + | ||
| - | function (Venkitasubramanian et al, 2006).</p> | + | |
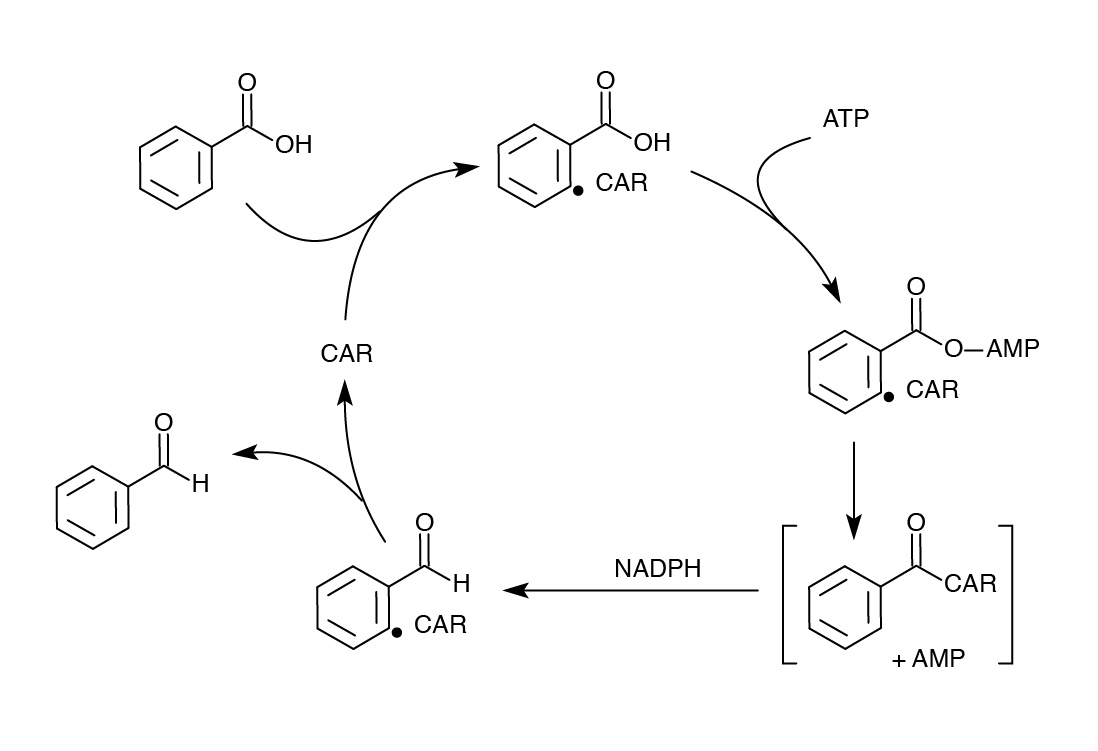
</html>[[File:Ucalgary Decarboxylation Team CAR Mechanism.jpg|center|350px|thumb|Figure 6. Mechanism of action of CAR]]<html> | </html>[[File:Ucalgary Decarboxylation Team CAR Mechanism.jpg|center|350px|thumb|Figure 6. Mechanism of action of CAR]]<html> | ||
| - | <p>Another enzyme with the potential to remove carboxyl groups from NAs is olefin-forming | + | <p>Another enzyme with the potential to remove carboxyl groups from NAs is olefin-forming fatty acid decarboxylase (<i>OleT</i>) from <i>Jeotgalicoccus sp. ATCC 8456</i>. This is a decarboxylase of the cytochrome P450 family that acts on fatty acids, but has also been documented to have low substrate specificity (Rude et al, 2011). What was attractive with this was that it was one single enzyme that go do the job of the PetroBrick! Now that we knew that our decarboxylation approach was valid, it was time to start testing and comparing this gene to the PetroBrick.</p> |
| - | + | <h2> Progress so far </h2> | |
| - | + | <p> <i>CAR</i> and <i>NPT</i> were cloned from the host organism <i>N. iowensis</i> (NRRL 5646). CAR was ligated into the PET vector and verified by a restriction digest while <i>NPT</i> was cloned into pSB1C3(<a | |
| - | + | href="http://partsregistry.org/wiki/index.php?title=Part:BBa_K902061">BBa_K902061.</a>) and similarly verified.</p> | |
| - | + | <p><i>CAR</i> was cloned into pET47b+ plasmid due to six illegal cut sites(one XbaI site, two EcoRI sites, and three NotI | |
| - | + | sites) which made it unsuitable for the BioBrick construction vectors. We first attempted to use a multi-site | |
| - | the | + | mutagenesis derived from the QuikChange® Multi Site Directed Mutagenesis Kit, but this showed little success. Instead, a more |
| - | was | + | time-consuming but effective series of conventional single-site <a href="https://2012.igem.org/Team:Calgary/Notebook/Protocols/mutagenesis">mutagenesis procedure</a> was favoured, using the Kappa Hi-Fi polymerase. The XbaI and EcoRI sites were eliminated |
| - | + | first so that <i>CAR </i> can be moved from the pET Vector and ligated into the PSB1C3 vector (<a href="http://partsregistry.org/wiki/index.php?title=Part:BBa_K902062">BBa_K902062.</a>). <i>OleT</i> was successfully amplified from the <i>Jeotgalicoccus sp. ATCC 8456</i>.<p> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | into the PSB1C3 vector (<a | + | |
| - | + | ||
| - | href="http://partsregistry.org/wiki/index.php? | + | |
| - | + | ||
| - | title=Part:BBa_K902062">BBa_K902062.</a>). OleT was successfully | + | |
| - | + | ||
| - | amplified from the <i>Jeotgalicoccus sp. ATCC 8456</i>.<p> | + | |
</p> | </p> | ||
| - | <p>Like CAR, | + | <p>Like <i>CAR</i>, <i>OleT</i> was inserted in a pET47b+ (Novagen) vector before placing it into a BioBrick vector, as two illegal cut sites adjacent to one another needed to be mutagenized. This part is now being ligated into pSB1C3. |
| - | + | ||
| - | <i>OleT</i> was inserted in a pET47b+ (Novagen) vector before placing it into a | + | |
| - | + | ||
| - | BioBrick, as two illegal cut sites adjacent to one another needed | + | |
| - | + | ||
| - | to be mutagenized. This part is now being ligated into pSB1C3. | + | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | below.</p> | + | We are currently in the process of constructing all three parts under contorl of a <i>tetR </i> promoter and ribosomal binding site (<a href="http://partsregistry.org/Part:BBa_J13002">BBa_J13002</a>), and then constructing these composite parts together as outlined below.</p> |
<h2>Final testing constructs</h2> | <h2>Final testing constructs</h2> | ||
| - | <p>Final testing constructs are almost complete. These are illustrated in figure 7 and will allow us to compare the three different approaches. Unfortunately, as Washington only sent us the PetroBrick and not the two individual components, we will have to compare a combination of the PetroBrick and CAR/NPT to the PetroBrick alone and to <i> | + | <p>Final testing constructs are almost complete. These are illustrated in figure 7 and will allow us to compare the three different approaches. Unfortunately, as Washington only sent us the PetroBrick and not the two individual components, we will have to compare a combination of the PetroBrick and CAR/NPT to the PetroBrick alone and to <i>OleT</i>. </p> |
<p></html>[[File:Ucalgary_Decarboxylation_Team_J13002+car+J13002+npt+PetroBrick.png|centre|450px]]<html></html>[[File:Ucalgary Decarboxylation Team J13002+oleT.png|centre|350px|thumb|Figure 7. Final constructs required for validating and comparing different decarboxylation approaches]]<html></p> | <p></html>[[File:Ucalgary_Decarboxylation_Team_J13002+car+J13002+npt+PetroBrick.png|centre|450px]]<html></html>[[File:Ucalgary Decarboxylation Team J13002+oleT.png|centre|350px|thumb|Figure 7. Final constructs required for validating and comparing different decarboxylation approaches]]<html></p> | ||
Revision as of 21:09, 3 October 2012


Hello! iGEM Calgary's wiki functions best with Javascript enabled, especially for mobile devices. We recommend that you enable Javascript on your device for the best wiki-viewing experience. Thanks!
Decarboxylation

Why Decarboxylation?
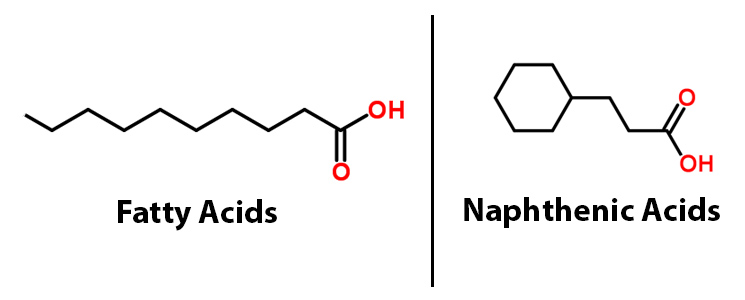
Among the toxins found in the tailing ponds, naphthenic acids (NAs) are among the most harmful and the most common. Though there is great diversity within the NAs class of compounds, all share the common chemical feature of a carboxylic acid group. The carboxyl group is the primary cause for their toxicity, allowing these chemicals to traverse cell membranes and react with cellular materials (Frank et al, 2009). NAs are recalcitranct (not easily degraded), potentially harmful to the surrounding ecosystem (Clemente & Fedorak, 2005) and corrosive to extraction and transport equipment of petroleum materials (Slavcheva et al, 1999). Corrosion of pipelines leads to higher maintenance costs as well as the grim possibility of these and other toxins leaking into the environment. There is a need for methods to degrade NAs that are not prohibitively expensive or that would result in production of other hazardous chemicals.
The main goal of OSCAR is to turn toxins like these into useable hydrocarbons by removing the carboxylic acid group(s) (Behar & Albrecht, 1984). Since NAs from petroleum deposits are a variable mixture, an enzymatic process with broad specificity is necessary. With the removal of the carboxylic acid moiety, we aim to produce alkanes suitable for use as fuel. The goal of this subproject was to find one or more suitable pathways to accomplish the decarboxylation of compounds such as NAs with the broadest specificity.
The PetroBrick
The 2011 Washington iGEM team developed the PetroBrick (BBa_K590025.), a BioBrick consisting of two primary genes. These include acyl-ACP reductase (AAR), which reduces fatty acids bound to ACP to fatty aldehydes, and a second gene called aldehyde decarbonylase (ADC), which subsequently cleaves the entire aldehyde group and results in a hydrocarbon chain. Essentially this allows for hydrocarbons to be produced from glucose. What we realized though, is that the fatty acids that the PetroBrick targets, have a very similar structure to NAs.
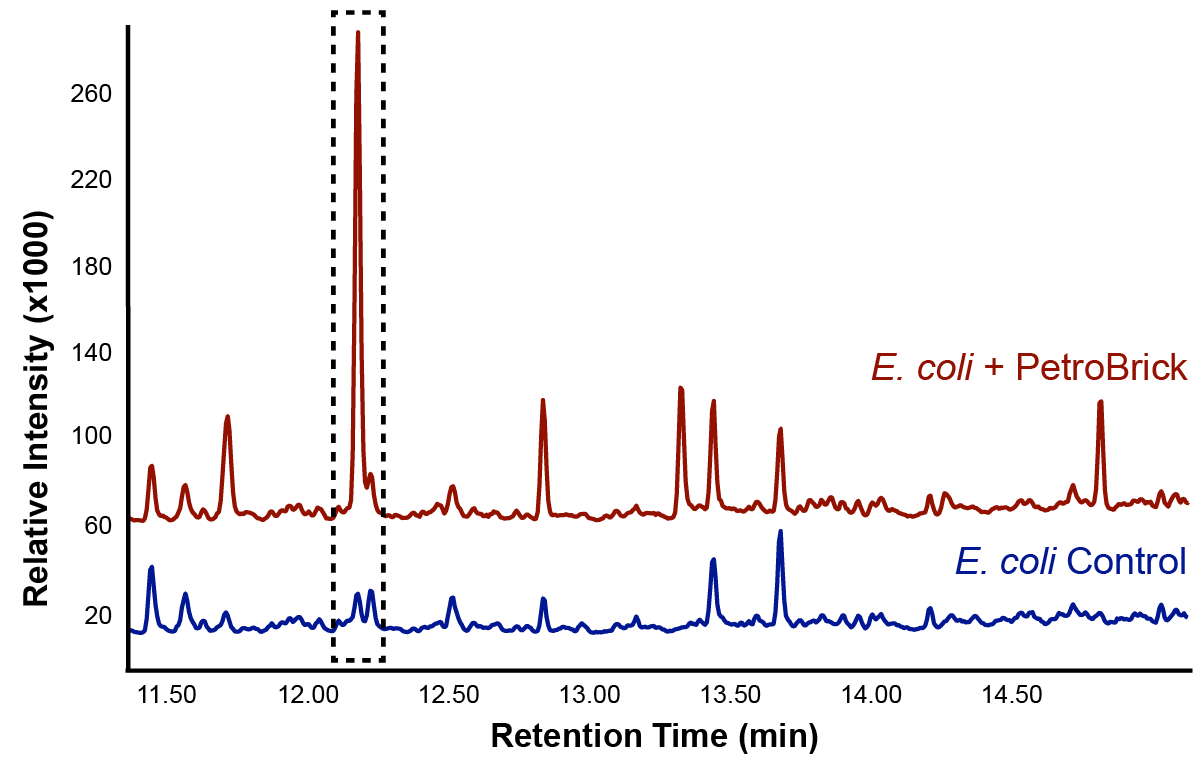
This lead us to believe that the PetroBrick may have the potential to turn NAs in to hydrocarbons and be a perfect solution to remediating NAs! First though, we needed to show that the PetroBrick did in fact work as expected. We had some difficulty with the DNA from the registry and had to request the constructs directly from the Washington team. Once we had the Petrobrick, we needed to verify that the Petrobrick would work in our hands as it did for the 2011 Washington team. Figures 2 and 3 demonstrate the function of the Petrobrick.
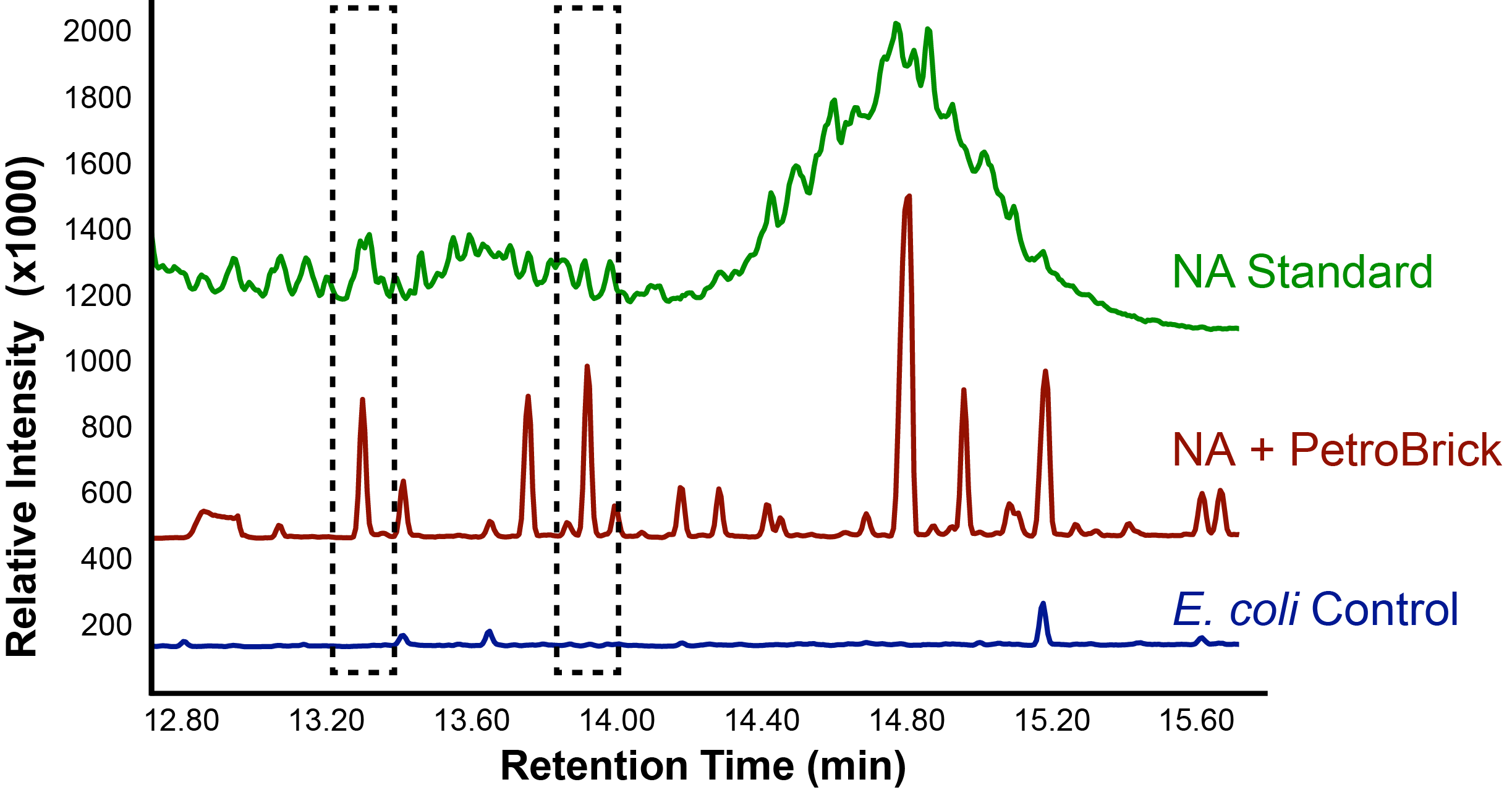
With the Petrobrick shown to be able to successfully produce alkanes, it was time to test it out on NAs, to see if they could be selectively converted into alkanes! This experiment used commercially available NAs fractions including a large number of different complex NAs compounds.
Successful conversion of NA's into Hydrocarbons!
The above graphs indicate that hydrocarbons were successfully produced from E.coli that contained the PetroBrick plasmid, as analysed with GC-MS. In Figure 2, E.coli containing the PetroBrick had significantly higher hydrocarbon peaks than in a control of E.coli that did not contain the PetroBrick plasmid. Not only was the PetroBrick able to degrade NAs into alkanes, but it was also able to produce alkenes as shown by Figure 3, indicating that the PetroBrick worked how we had expected it to!
Nocardia Carboxylic Acid Reductase (CAR)- Can we do better?
Although we were successful using the Petrobrick to remove carboxyl groups from NAs, we wanted to improve on our results to see if we could get a higher yield or possibly target other compounds. One of our original fears in using the PetroBrick to decarboyxlate NAs was that the first enzyme AAR was reported to be highly specific for fatty acids bound to ACP. We had concerns about its compatibility with NAs and therefore sought another enzyme in the literature called carboxylic acid reductase (CAR) that was documented to perform a similar task as AAR, converting fatty acids to aldehydes, but with much lower specificity (He et al, 2004). This enzyme, from N. iowensis does not require covalent attachment to ACP so would likely be much broader in substrate specificity. It requires a second gene from N. iowensis, called Nocardia phosphopantetheinyl transferase (NPT) necessary to append a 4’- phosphopantetheine prosthetic group to CAR required for its full function (Venkitasubramanian et al, 2006).
Another enzyme with the potential to remove carboxyl groups from NAs is olefin-forming fatty acid decarboxylase (OleT) from Jeotgalicoccus sp. ATCC 8456. This is a decarboxylase of the cytochrome P450 family that acts on fatty acids, but has also been documented to have low substrate specificity (Rude et al, 2011). What was attractive with this was that it was one single enzyme that go do the job of the PetroBrick! Now that we knew that our decarboxylation approach was valid, it was time to start testing and comparing this gene to the PetroBrick.
Progress so far
CAR and NPT were cloned from the host organism N. iowensis (NRRL 5646). CAR was ligated into the PET vector and verified by a restriction digest while NPT was cloned into pSB1C3(BBa_K902061.) and similarly verified.
CAR was cloned into pET47b+ plasmid due to six illegal cut sites(one XbaI site, two EcoRI sites, and three NotI sites) which made it unsuitable for the BioBrick construction vectors. We first attempted to use a multi-site mutagenesis derived from the QuikChange® Multi Site Directed Mutagenesis Kit, but this showed little success. Instead, a more time-consuming but effective series of conventional single-site mutagenesis procedure was favoured, using the Kappa Hi-Fi polymerase. The XbaI and EcoRI sites were eliminated first so that CAR can be moved from the pET Vector and ligated into the PSB1C3 vector (BBa_K902062.). OleT was successfully amplified from the Jeotgalicoccus sp. ATCC 8456.
Like CAR, OleT was inserted in a pET47b+ (Novagen) vector before placing it into a BioBrick vector, as two illegal cut sites adjacent to one another needed to be mutagenized. This part is now being ligated into pSB1C3. We are currently in the process of constructing all three parts under contorl of a tetR promoter and ribosomal binding site (BBa_J13002), and then constructing these composite parts together as outlined below.
Final testing constructs
Final testing constructs are almost complete. These are illustrated in figure 7 and will allow us to compare the three different approaches. Unfortunately, as Washington only sent us the PetroBrick and not the two individual components, we will have to compare a combination of the PetroBrick and CAR/NPT to the PetroBrick alone and to OleT.
 "
"