Team:Calgary/Project/HumanPractices/Killswitch/Regulation
From 2012.igem.org
Emily Hicks (Talk | contribs) |
|||
| Line 1: | Line 1: | ||
{{Team:Calgary/TemplateProjectOrange| | {{Team:Calgary/TemplateProjectOrange| | ||
| - | TITLE=Regulation/Expression Platform| | + | TITLE=Regulation/Expression Platform| |
| - | <html> | + | CONTENT=<html> |
<img src="https://static.igem.org/mediawiki/2012/8/8c/UCalgary2012_FRED_Killswitch_Regulation_Low-Res.png" style="float: right; padding: 10px; height: 200px;"></img> | <img src="https://static.igem.org/mediawiki/2012/8/8c/UCalgary2012_FRED_Killswitch_Regulation_Low-Res.png" style="float: right; padding: 10px; height: 200px;"></img> | ||
Revision as of 10:58, 3 October 2012
{{Team:Calgary/TemplateProjectOrange| TITLE=Regulation/Expression Platform|
CONTENT=
Tight Regulation
Inducible kill systems are not new to iGEM. Looking through the registry, there are several constructs such as the inducible bamHI system contributed by Berkley in 2006 (BBa_I716462) and tested by Lethbridge in 2011. This uses a bamHI gene downsteam of an arabinose-inducible promoter. Another example is an IPTG inducible Colicin construct (BBa_K117009) submitted by NTU-Singapore in 2008. One major problem with these systems however is a lack of tight control. As was demonstrated by the Lethbridge team, this part has leaky expression when uninduced. The frequently used lacI promoter has similar prblems when not used in conjunction with strong plasmid-medated expresson of lacI. This can be seen in our electrochemical characterization of the uidA hydrolase enzyme shown here. Tight control is not only a problem for kill switch application, but for any application requiring strict regulation. As such, we decided that expanding the registry repertoire of control elements would be useful for our system as well as a variety of other applications
Introducing the Riboswitch
Riboswitches are small pieces of mRNA that can bind to small molecules, essentially preventing translation of downstream from occurring. Thus by putting these in front of a gene of interest, we can regulate expression. This allows us another level of control on top of transcriptional regulation from promoter elements. Thus a combination of promoters and riboswitches could lead to much tighter control. This could present an easy solution to our problem. As there are many riboswitches that bind inexpensive ions, these could be cheap and feasible solution for our system
We explored 3 diffeent riboswitches, each responsive to a different metabolite: magneisum, manganese and molybdate that would be cheap to implement into a bioreactor environment. In addition, we also investigated an inducible/repressible promoter repsonvie to glucose and rhmnose
Magnesium repressible system
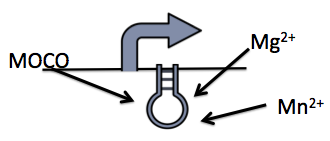
This system is repressed in the presence of magnesium. This system has two control components – a promoter and a riboswitch. Normally the magnesium promoter (mgtA promoter) and the magnesium riboswitch (mgtArb) are activated if there is a deficiency of magnesium in the cell. The lack of magnesium activates other genes in E. coli to allow influx of magnesium into the cell. There are two proteins in the cascade that activate the system namely PhoP and PhoQ. PhoQ is the trans-membrane protein which gets activated in the absence of magnesium and phosphorylates PhoP. PhoP in turn binds to the mgtA promoter and transcribes genes downstream.
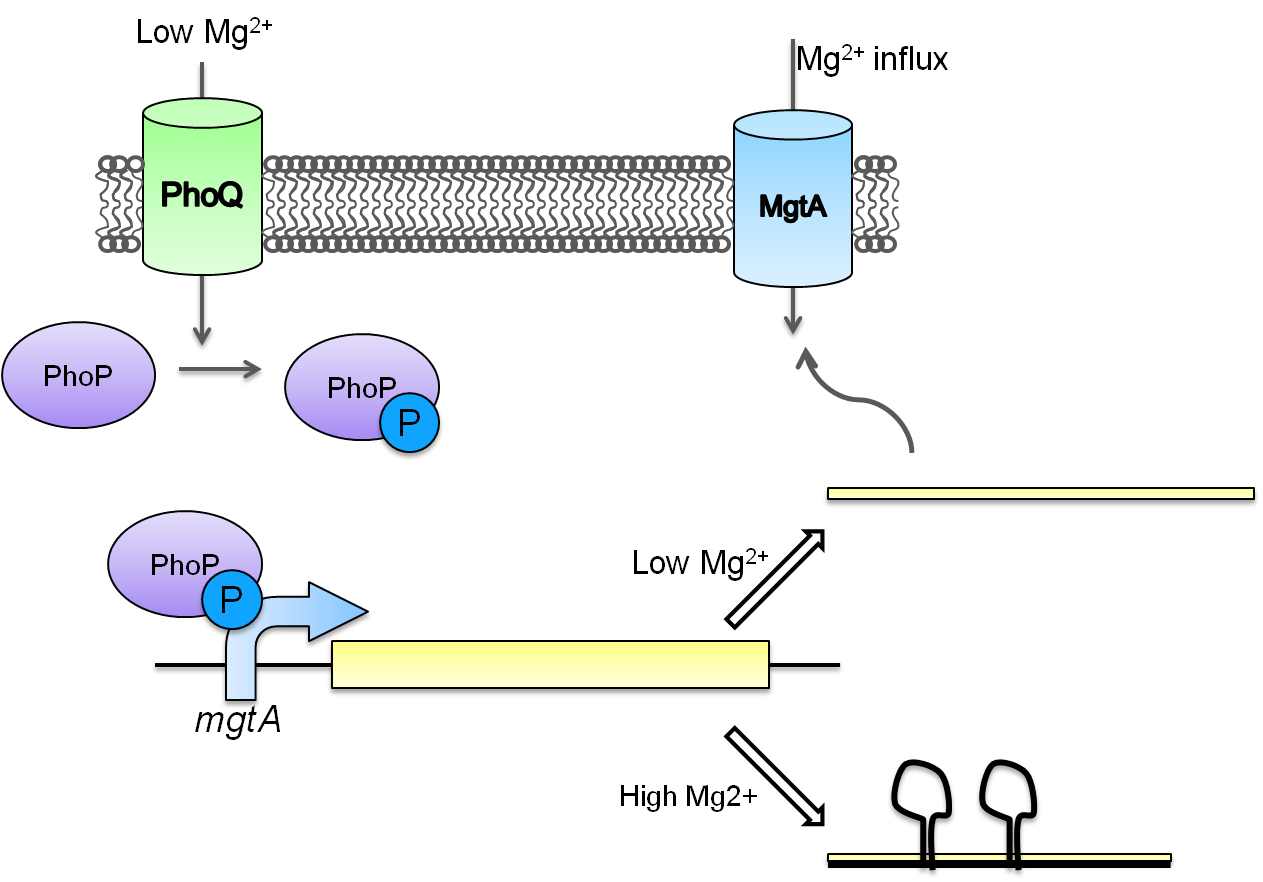
Test circuits for the magnesium system
To test the magnesium regulatory elements we built each of the elements with a reporter gene. We chose Bba_K082003 which is GFP with an LVA tag as our choice of reporter. We did not choose BBa_E0040, the stable GFP, because we wanted a real time indication of the system's control. Stable GFP has a half life of 8 hours and would still fluoresce when the system is shut off.
We build these circuits to test the control elements of the system, namely the mgtA promoter and the mgtA riboswitch.
Characterization of these circuits
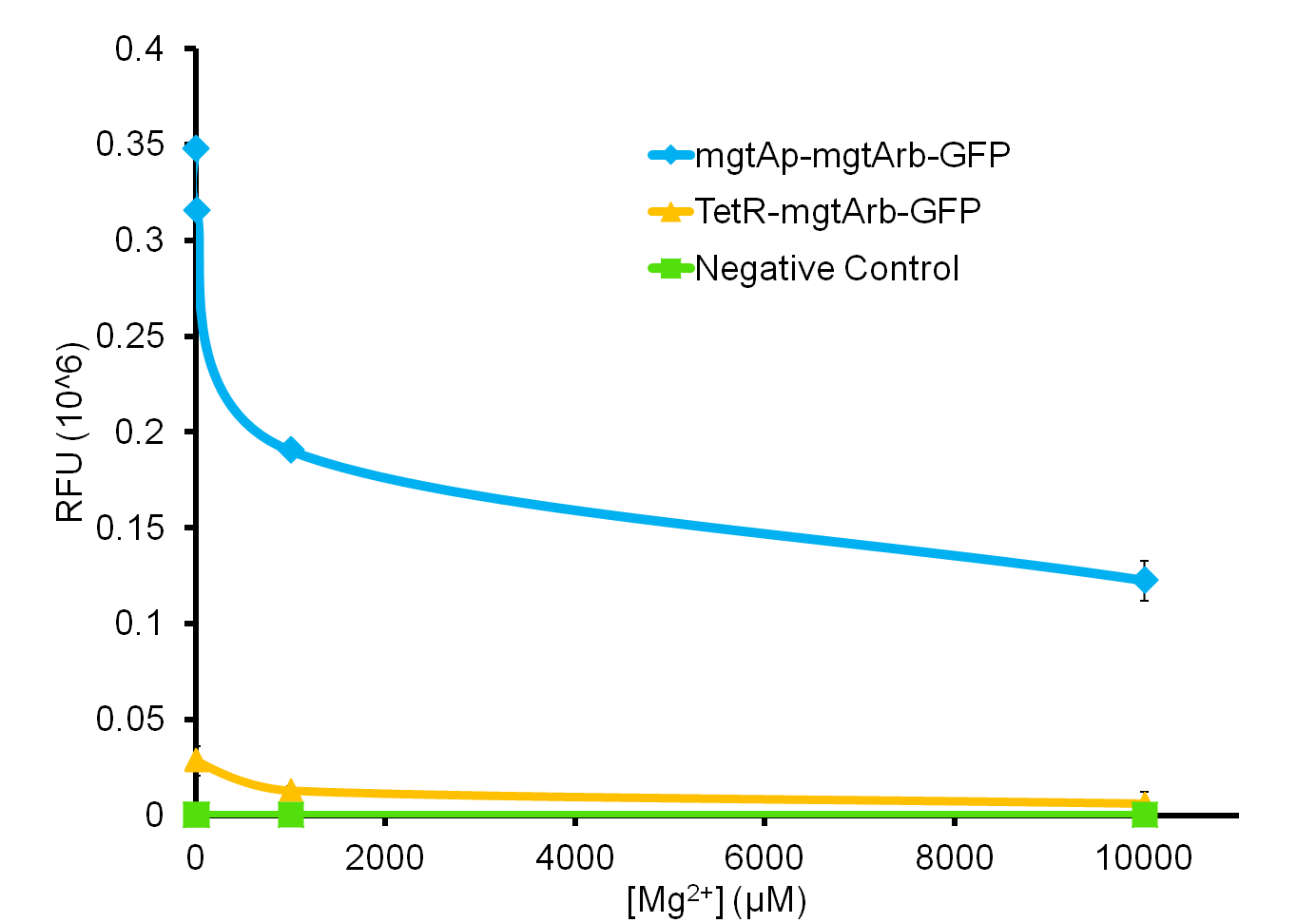
We tested the aforementioned circuits in different concentrations of magnesium. For detailed protocol see INSERT LINK HERE. The values were normalized to the negative control which is the magnesium promoter and riboswitch alone.
There is a much larger decrease in the GFP output when the mgtA promoter and riboswitch are working together compared to the mgtA riboswitch alone under the control of TetR promoter. This suggests that having both the promoter and the riboswitch together provides a tighter control over the genes expressed downstream. This also suggests that magnesium riboswitch alone is sufficient in reducing gene expression downstream of a constitutive promoter.
It is important to consider however that the control elements of the system namely PhoP and PhoQ were not present in the circuits tested and therefore there is some GFP expression in even at the inhibitory concentration (10mM MgCl2). We believe that having the regulatory elements would give us better control and get rid of the leakiness.
Although the magnesium system is highly regulated, it is not a suitable system for the purposes of our bioreactor. The tailings are composed of very high concentration of magnesium- upto 120mM(Kim et al, 2011). As can be seen from figure 3, this would inhibit the system. Therefore, if our bacteria escapes into the tailings, the kill genes would not be activated and the bacteria would be able to survive.
In contrast, it is important to note that this system adds important regulatory elements to the registry such as an inducible promoter and a riboswitch which can be used by other teams to control both killswitches as well as other regulatory pathways which do not pertain using tailings.
We also wanted to test the magnesium system with our kill genes. The micrococcal nuclease that arrived from IDT had 1bp mutation which changed a isoleucine residue into a lysine. Therefore we had to mutate it. To test the circuits we measured both CFU and OD 600.
[[File:24 hour assay with mgtap-rb-S7 Ucalgary.pn
 "
"