Team:Calgary/Project/HumanPractices/Killswitch/Regulation
From 2012.igem.org
| Line 1: | Line 1: | ||
{{Team:Calgary/TemplateProjectOrange| | {{Team:Calgary/TemplateProjectOrange| | ||
TITLE=Regulation/Expression Platform| | TITLE=Regulation/Expression Platform| | ||
| - | + | CONTENT= | |
| - | CONTENT=<html> | + | <html> |
<img src="https://static.igem.org/mediawiki/2012/8/8c/UCalgary2012_FRED_Killswitch_Regulation_Low-Res.png" style="float: right; padding: 10px; height: 200px;"></img> | <img src="https://static.igem.org/mediawiki/2012/8/8c/UCalgary2012_FRED_Killswitch_Regulation_Low-Res.png" style="float: right; padding: 10px; height: 200px;"></img> | ||
| Line 17: | Line 17: | ||
</html> [[File:UofC_RIBOSWITCH.png|centre|350px]] <html> | </html> [[File:UofC_RIBOSWITCH.png|centre|350px]] <html> | ||
| - | <p>We explored 3 diffeent riboswitches, each responsive to a different metabolite | + | <p>We explored 3 diffeent riboswitches, each responsive to a different metabolite that would be cheap to implement into a bioreactor environment. In addition, we also investigated an inducible/repressible promoter repsonvie to glucose and rhmnose</p> |
| Line 50: | Line 50: | ||
<p> We also wanted to test the magnesium system with our kill genes. The micrococcal nuclease that arrived from IDT had 1bp mutation which changed a isoleucine residue into a lysine. Therefore we had to mutate it. To test the circuits we measured both CFU and OD 600.</p> | <p> We also wanted to test the magnesium system with our kill genes. The micrococcal nuclease that arrived from IDT had 1bp mutation which changed a isoleucine residue into a lysine. Therefore we had to mutate it. To test the circuits we measured both CFU and OD 600.</p> | ||
| - | </html>[[File:24 hour assay with mgtap-rb-S7 Ucalgary. | + | </html>[[File:24 hour assay with mgtap-rb-S7 Ucalgary.png|thumb|750px|center| Figure 4: This shows the OD600 values of mgtA circuits with S7 both mutated and unmutated. The negative control consists of <i>mgtAp-mgtArb</i>.]]<html> |
| + | <p> Figure 4 shows that the mgtAp-mgtArb-S7 starts acting approximately 4 hours after induction. However, it also shows that 10mM MgCl<sub>2</sub> is not enough salt to inhibit the entire system because there is no difference in OD600 measurement at 4hr timepoint between 10mM and the 0mM concentrations. This test needs to be repeated with higher concentrations of Mg<sup>2+</sup> however this data suggests that the mutagenesis was successful and S7 is active and killing the cells at approximately 4hr which does not necessarily reflect upon the activity of S7 but also reflects upon the response time of the mgtA system.</p> | ||
| + | |||
| + | <h1>Manganese regulation</h1> | ||
| + | |||
| + | <p> Manganese is an essential micronutrient. It is an important co-factor for enzymes and it also reduces oxidative stress in the cell(Waters et al, 2011). Despite being an important micronutrient it is toxic to cells at high levels. MntR detects the level of manganese in the cell and binds acts as a transcription factor to control the expression of manganese transportes such as mntH, mntP and mntABCDE. In order to regulate these genes mntR binds to the mntP promoter. The manganese homeostatsis is also controlled by the manganese riboswitch <i>mntPrb</i></p> | ||
| + | |||
| + | </html>[[File:U.Calgary.2012_10.02.2012_Membrane_Final_2.png|thumb|500px|center|Figure 4: The manganese system in the presence of manganese found in the tailing ponds will initially trigger the mntR regulator. As one of its function, the regulator will repress the mntH transporter preventing the movement of manganese. As a second function, the mntR will upregulate the putative efflux pump. The manganese system itself responds to the manganese metal allowing the transcription of the gene downstream. The addition of the mntR in this system is generally used to better regulate the manganese system.]]<html> | ||
| + | |||
| + | <h3><i>Our manganese system</i></h3> | ||
| + | |||
| + | <p>For our system we use the mntR,<i>mntP</i> promoter and <i>mntP</i> riboswitch. At high levels of manganese the mntP promoter <i>mntPp</i> and riboswitch <i>mntPrb</i> is activated producing the genes downstream. For our purposes we will be constructing <i>mntPp</i> and <i>mntPrb</i> with S7 and CviAII. </p> | ||
| + | |||
| + | </html>[[File:U.Calgary.2012_10.02.2012_Final_Construct_1.png|thumb|600px|center|Figure 5: Final construct for the manganese system. The circuit includes a TetR promoter, RBS, mntR, double terminator, <i>mntP promoter</i>, <i>mntP riboswitch</i>, S7, <i>mntP riboswitch</i> and CViAII.]]<html> | ||
| + | |||
| + | </html>[[File:U.Calgary.2012_10.02.2012_Construct_3.png|thumb|200px|right|Figure 6: These set of circuits are used to characterise both the mntP promoter and the mntP riboswitch. The negative control composes of mntP promoter-mntP riboswitch while the positive control is the circuit TetR-RBS-K082003.]]<html> | ||
| + | |||
| + | <h3><i>Test circuits for the manganese system</i></h3> | ||
| + | |||
| + | <p>The manganese system was constructed with GFP-LVA for characterization purposes. The following are the control circuits built in order to characterise the MntP promoter and the MntP riboswitch.</p> | ||
| + | |||
| + | |||
| + | <h3><i>Future Use</i></h3> | ||
| + | |||
| + | <p>Even though this system is controlled and a good way to regulate kill genes, there are some limitations to this system. Tailings water contain high levels of Mn2+ in the contaminated water ~48mg/L or ~170µM/L. This concentration is 17 times the amount of Mn2+ needed to trigger the system (10 µM) therefore an additional chemical such as EDTA (a chelator) will have to be added to lower the manganese levels in the bioreactor. This however requires us to add EDTA into each set of tailings that is added into the bioreactor. EDTA is expensive and will thereby increase the cost of our bioreactor. Although this regulation method may not be feasible for our bioreactor, this regulatory system may be used in another pathway.</p> | ||
| + | |||
| + | |||
| + | <a name="rhamnose"></a> | ||
| + | <h1>Glucose repressible system</h1> | ||
| + | |||
| + | <h2>Purpose</h2> | ||
| + | <p>The rhamnose inducible promoter is suitable for controlling the kill switch in the the bioreactor since the promoter is shown to be tightly repressed with glucose. We aim to supplement the bioreactor with glucose to inhibit | ||
| + | expression of the kill genes in the bioreactor. Escape of bacteria into the tailings ponds will cause expression of the kill genes since glucose levels in the surrounding environment would be insufficient for deactivate the promoter. We selected this system for two reasons. Firstly, the promoter exhibits tight repression with glucose and could thus limit inadvertent expression of the kill genes; and secondly, glucose is a common monosacharide which is economically feasible to continuously supplement into the bioreactor. | ||
| + | </p> | ||
| + | |||
| + | <h2>Application of the rhamnose promoter</h2> | ||
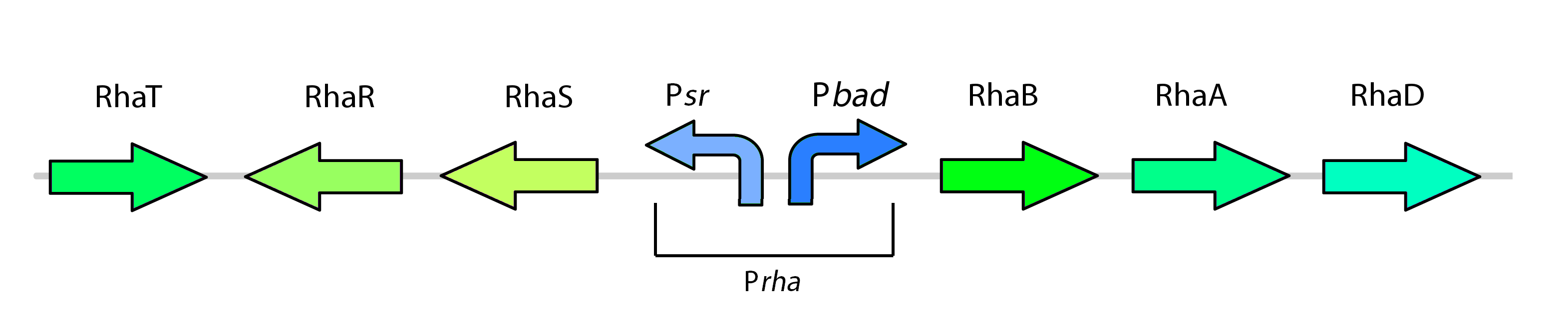
| + | <p>The rhamnose promoter (pRha) is responsible for regulating six genes related to rhamnose metabolism and contains a separate promoter on its leading and reverse strands (see Figure one). RhaR and RhaS serve to regulate expression of the rhamnose metabolism genes rhaB, rhaA, and rhaD on the opposite side of the promoter. The RhaR transcription factor is activated by L-rhamnose to up-regulate expression rhaSR operon. In turn, the resulting RhaS activates the rhaBAD operon to generate the rhamnose metabolism genes (Egan & Schleif, 1993).</p> | ||
| + | |||
| + | </html>[[File:NativeRhamnosePromoter_Calgary2012.jpg|thumb|500px|center|Figure one: The rhamnose metabolism genes as they exist in Top Ten E. coli]] | ||
<html> | <html> | ||
| + | |||
| + | <p>As a kill switch regulator, our team has harnessed global catabolite repression of the rhamnose promoter. Expression of the rhaBAD operon with RhaS requires the binding of catabolite receptor protein (CRP) cAMP complex to the promoter. When glucose is present, cAMP levels are low, such that CRP is unable to activate the promoter (Egan & Schleif, 1993). With this mechanism, our team set out to control our kill genes with glucose with the following rhamnose promoter construct:</p> | ||
| + | |||
| + | </html>[[File:RhamnoseConstruct_Calgary2012.png|thumb|500px|center|Figure two: Our design for the glucose repressible, rhamnose based promoter for our kill system.]]<html> | ||
| + | |||
| + | <p>Our kill system is different from the native rhamnose system with the rhaR and rhaS control genes. We have constituitively expressed rhaS to overcome dependency on rhamnose to cause activation of the kill switch. While RhaS is continuously present, global catabolite repression prevents its activation the kill genes in the bioreactor; in the outside environment, glucose levels are lower such that RhaS is able to activate the kill genes.</p> | ||
| + | |||
| + | <h2>Components and characterization</h2> | ||
| + | <p>Our team had pRha promoter commercially synthesized as per the sequence given by Jeske and Altenbuchner (2010). The rhaS and rhaR genes were amplified via PCR from Top Ten E. coli using Kapa Hi-Fi polymerase. </p> | ||
| + | |||
| + | <p>Given the control gene modifications which we have engineered into our system to optimize it for the tailings ponds, we are working to determine whether glucose repression of our modified system match patterns shown by Giacalone et al. (2006) and Jeske and Altenbuchner (2010). To this end, we have constructed the rhamnose promoter with GFP and are finalizing construction with constituitively expressed RhaS, so that we can characterize (LINK) this promoter and test it with our kill genes.</p> | ||
| + | |||
| + | |||
| + | |||
| + | <h1>The Moco Riboswitch </h1> | ||
| + | <h2>Background</h2> | ||
| + | <p> | ||
| + | The molybdenum cofactor (moco) riboswitch is an RNA element which responds to the presence of the metabolite molybdenum cofactor (Regulski et al, 2008). This RNA element is located in the E.coli genome just upstream of the moaABCDE operon, which contain the important moco synthesis genes. Moco is an important cofactor in many different enzymes ranging from to this. The moco riboswitch has 2 regions: an aptamer domain and the expression platform. When moco is present in the cell it will bind to the aptamer region in the riboswitch which will cause an allosteric change. This allosteric change affects the expression platform by physically hiding the ribosome binding site thus preventing translation from occurring and hence adding a translational level of gene expression regulation. Therefore, the moco riboswitch, when activated by moco, inhibits gene expression. | ||
| + | |||
| + | </p> | ||
| + | </html> | ||
| + | [[File:Moco_riboswitchCalgary2012.jpg|thumb|500px|center|Figure 9: This picture depicts the Moco RNA motif which is upstream of the moaABCDE operon. ]]<html> | ||
| + | |||
| + | <h2>The Killswitch Design</h2> | ||
| + | |||
| + | <p> | ||
| + | This riboswitch system can be used to design a killswitch mechanism to regulate the expression of our kill genes, CviAII and S7. The basic design of this system includes the kill genes downstream of the riboswitch all of which is under the control of a constitutive promoter. Downstream of this construct is the moa operon constitutively expressed. This entire construct will be present in a low copy plasmid in our bacteria. While moco synthesis is a normal process in the bacteria we wanted to constitutively express the moa operon to up regulate the expression of moco. This is to ensure high enough concentrations of moco present in order to inactivate the kill switch when the bacteria are in the bioreactor. | ||
| + | |||
| + | </p> | ||
| + | |||
| + | <h2>Molybdate in TPW</h2> | ||
| + | <p> | ||
| + | |||
| + | Molybdenum, Mo, is a trace element that is required by the bacteria for moco synthesis. Bacteria cannot uptake molybdenum in the elemental form and so uptakes molybdenum in its oxyanion form molybdate, MoO4, using the molybdate transport system. We wanted a system where molybdate is present in the bioreactor permitting moco synthesis (inactivate killswitch) and absent in the tailings pond water (TPW) preventing moco synthesis (activate killswitch). We discovered that molybdenum is indeed present in the TPW at around 1 µM. </p> | ||
| + | <p> | ||
| + | Upon further literature research we found that molybdate is transported using a molybdate transport system which is coded by the modABCD operon. It has been shown that knocking out modC enables the transport system dysfunctional and prevents moco synthesis. However, if molybdate is supplemented in high enough concentrations to the bacteria, the bacteria are able to use its sulphate transport system to transport the molybdate in. This gave rise to the idea of having a modC knock out strain of bacteria in our system and supplementing it with molybdate. The paper tried 10mM sodium molybdate supplementation in the media to repress gene expression whereas that the literature level states that there is less than 1 µM molybdate in the TPW. Therefore, in the tailings ponds, the concentration of molybdate is not high enough to be transported into the cell using the sulphate transport mechanism. This prevents moco synthesis and activates the killswitch. While sodium molybdate is expensive and can be argued to be impractical to provide on a large bioreactor scale, there are many inexpensive and effective filtration mechanisms available to filter out most of the molybdate and reuse it. </p> | ||
| + | |||
| + | <h2>Characterization</h2> | ||
| + | |||
| + | <p>There are two experiments that we want to run with this system. First, we wish to characterize the concentration of molybdate needed in the system to enter through the sulphate transport mechanism. We will supply lab strain E.Coli and modC knockout strain of E.Coli with varying concentrations of molybdate and measure the optical density. This will be followed by a CFU assay. This experiment will also inform us of toxic levels of molybdate for our bacteria. Our second experiment is the characterization of the moco killswitch system. </p> | ||
| + | |||
| + | <h2> Final Moco Construct: </h2> | ||
| + | |||
| + | </html> | ||
| + | [[File:Final_Moco_ConnstructCalgary2012.jpg|thumb|500px|center|Figure 10: This is the final moco riboswitch construct. The riboswitch regulated kill genes will be constitutively expressed ensuring that the kill genes are activated should the bacteria escape the system. The moaABCDE operon is constitutively expressed so that moco synthesis is up regulated when there is molybdate in the system.]]<html> | ||
| + | |||
| + | </html> | ||
| + | }} | ||
Revision as of 10:59, 3 October 2012


Hello! iGEM Calgary's wiki functions best with Javascript enabled, especially for mobile devices. We recommend that you enable Javascript on your device for the best wiki-viewing experience. Thanks!
Regulation/Expression Platform

Tight Regulation
Inducible kill systems are not new to iGEM. Looking through the registry, there are several constructs such as the inducible bamHI system contributed by Berkley in 2006 (BBa_I716462) and tested by Lethbridge in 2011. This uses a bamHI gene downsteam of an arabinose-inducible promoter. Another example is an IPTG inducible Colicin construct (BBa_K117009) submitted by NTU-Singapore in 2008. One major problem with these systems however is a lack of tight control. As was demonstrated by the Lethbridge team, this part has leaky expression when uninduced. The frequently used lacI promoter has similar prblems when not used in conjunction with strong plasmid-medated expresson of lacI. This can be seen in our electrochemical characterization of the uidA hydrolase enzyme shown here. Tight control is not only a problem for kill switch application, but for any application requiring strict regulation. As such, we decided that expanding the registry repertoire of control elements would be useful for our system as well as a variety of other applications
Introducing the Riboswitch
Riboswitches are small pieces of mRNA that can bind to small molecules, essentially preventing translation of downstream from occurring. Thus by putting these in front of a gene of interest, we can regulate expression. This allows us another level of control on top of transcriptional regulation from promoter elements. Thus a combination of promoters and riboswitches could lead to much tighter control. This could present an easy solution to our problem. As there are many riboswitches that bind inexpensive ions, these could be cheap and feasible solution for our system
We explored 3 diffeent riboswitches, each responsive to a different metabolite that would be cheap to implement into a bioreactor environment. In addition, we also investigated an inducible/repressible promoter repsonvie to glucose and rhmnose
Magnesium repressible system
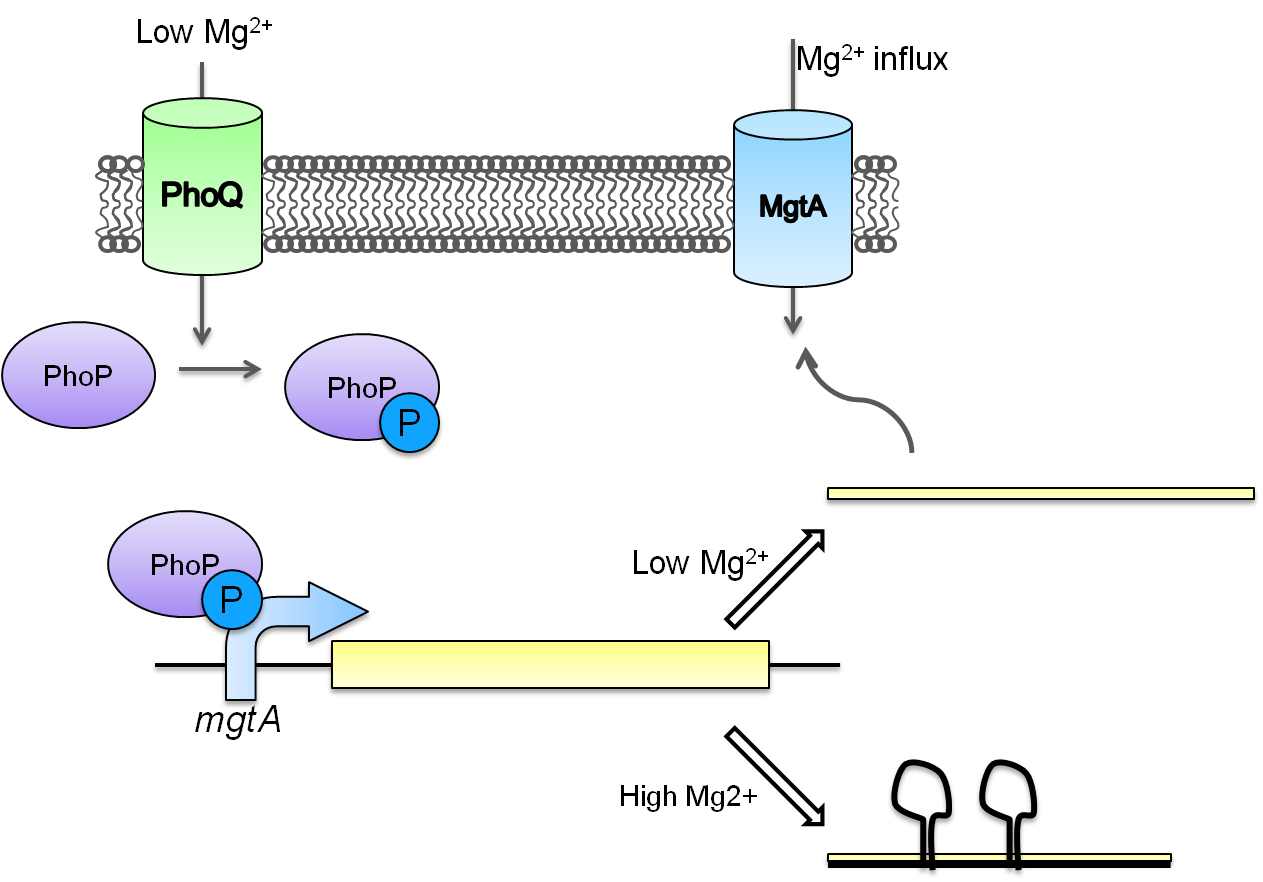
This system is repressed in the presence of magnesium. This system has two control components – a promoter and a riboswitch. Normally the magnesium promoter (mgtA promoter) and the magnesium riboswitch (mgtArb) are activated if there is a deficiency of magnesium in the cell. The lack of magnesium activates other genes in E. coli to allow influx of magnesium into the cell. There are two proteins in the cascade that activate the system namely PhoP and PhoQ. PhoQ is the trans-membrane protein which gets activated in the absence of magnesium and phosphorylates PhoP. PhoP in turn binds to the mgtA promoter and transcribes genes downstream.
Test circuits for the magnesium system
To test the magnesium regulatory elements we built each of the elements with a reporter gene. We chose Bba_K082003 which is GFP with an LVA tag as our choice of reporter. We did not choose BBa_E0040, the stable GFP, because we wanted a real time indication of the system's control. Stable GFP has a half life of 8 hours and would still fluoresce when the system is shut off.
We build these circuits to test the control elements of the system, namely the mgtA promoter and the mgtA riboswitch.
Characterization of these circuits
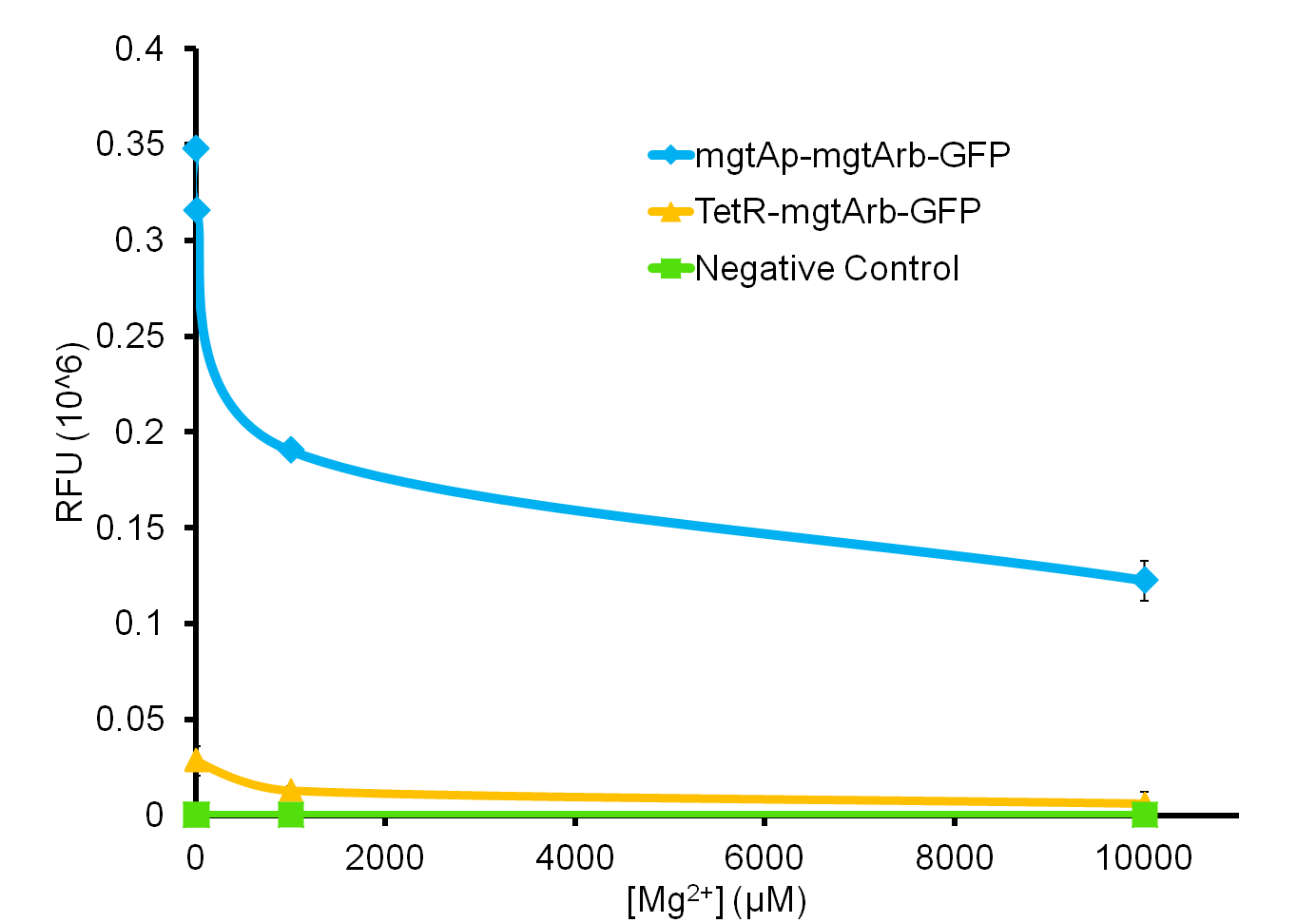
We tested the aforementioned circuits in different concentrations of magnesium. For detailed protocol see INSERT LINK HERE. The values were normalized to the negative control which is the magnesium promoter and riboswitch alone.
There is a much larger decrease in the GFP output when the mgtA promoter and riboswitch are working together compared to the mgtA riboswitch alone under the control of TetR promoter. This suggests that having both the promoter and the riboswitch together provides a tighter control over the genes expressed downstream. This also suggests that magnesium riboswitch alone is sufficient in reducing gene expression downstream of a constitutive promoter.
It is important to consider however that the control elements of the system namely PhoP and PhoQ were not present in the circuits tested and therefore there is some GFP expression in even at the inhibitory concentration (10mM MgCl2). We believe that having the regulatory elements would give us better control and get rid of the leakiness.
Although the magnesium system is highly regulated, it is not a suitable system for the purposes of our bioreactor. The tailings are composed of very high concentration of magnesium- upto 120mM(Kim et al, 2011). As can be seen from figure 3, this would inhibit the system. Therefore, if our bacteria escapes into the tailings, the kill genes would not be activated and the bacteria would be able to survive.
In contrast, it is important to note that this system adds important regulatory elements to the registry such as an inducible promoter and a riboswitch which can be used by other teams to control both killswitches as well as other regulatory pathways which do not pertain using tailings.
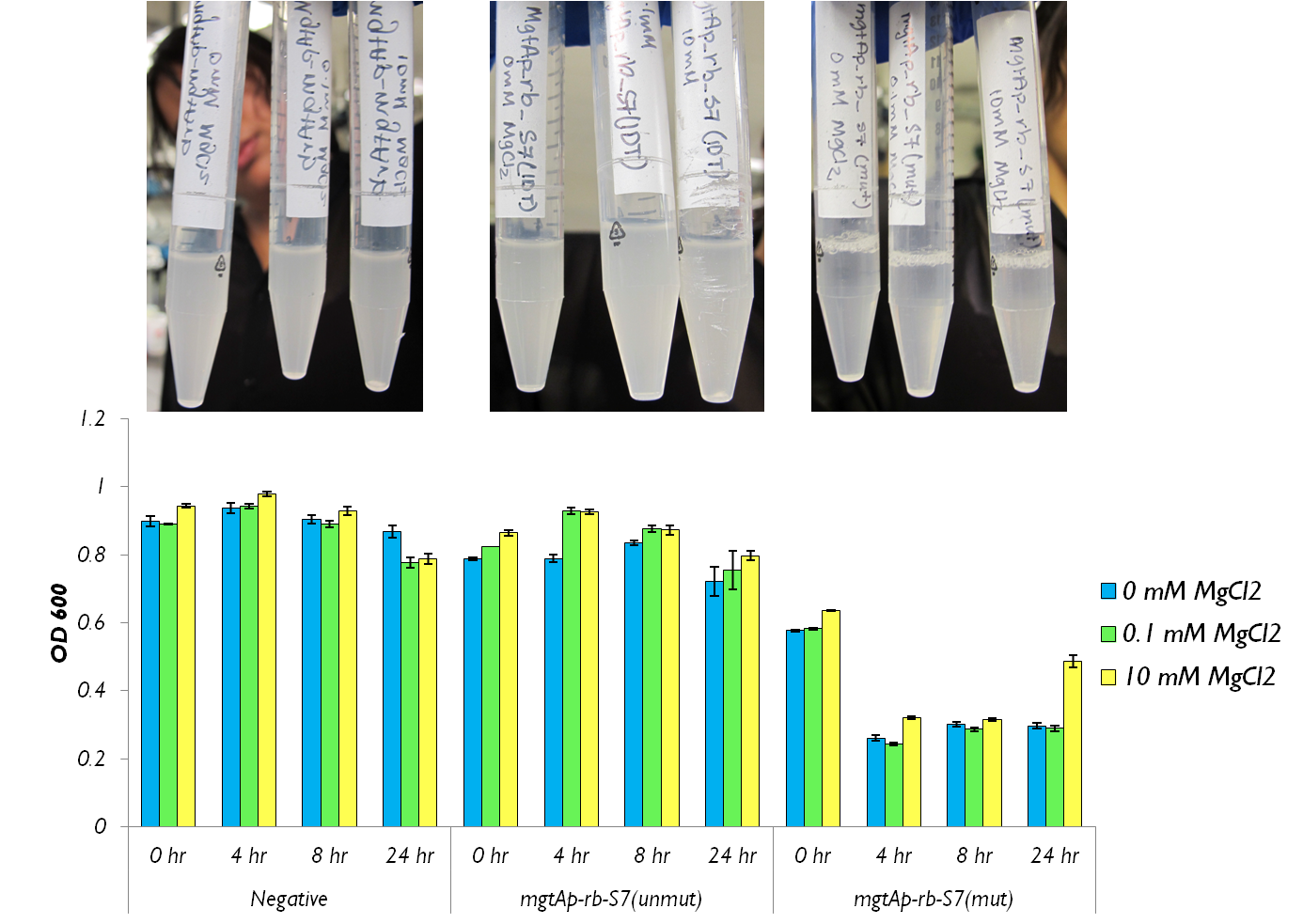
We also wanted to test the magnesium system with our kill genes. The micrococcal nuclease that arrived from IDT had 1bp mutation which changed a isoleucine residue into a lysine. Therefore we had to mutate it. To test the circuits we measured both CFU and OD 600.
Figure 4 shows that the mgtAp-mgtArb-S7 starts acting approximately 4 hours after induction. However, it also shows that 10mM MgCl2 is not enough salt to inhibit the entire system because there is no difference in OD600 measurement at 4hr timepoint between 10mM and the 0mM concentrations. This test needs to be repeated with higher concentrations of Mg2+ however this data suggests that the mutagenesis was successful and S7 is active and killing the cells at approximately 4hr which does not necessarily reflect upon the activity of S7 but also reflects upon the response time of the mgtA system.
Manganese regulation
Manganese is an essential micronutrient. It is an important co-factor for enzymes and it also reduces oxidative stress in the cell(Waters et al, 2011). Despite being an important micronutrient it is toxic to cells at high levels. MntR detects the level of manganese in the cell and binds acts as a transcription factor to control the expression of manganese transportes such as mntH, mntP and mntABCDE. In order to regulate these genes mntR binds to the mntP promoter. The manganese homeostatsis is also controlled by the manganese riboswitch mntPrb
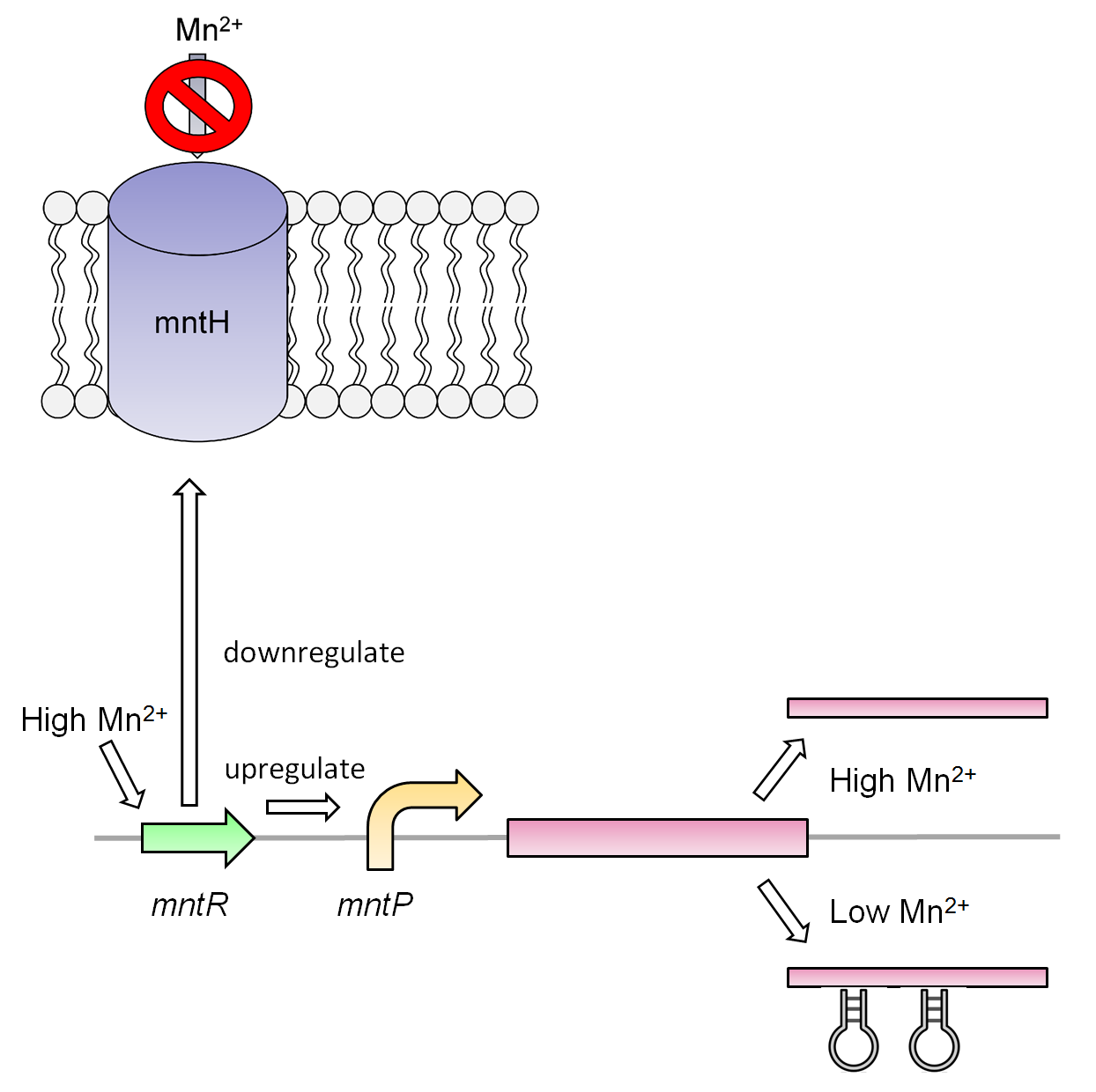
Our manganese system
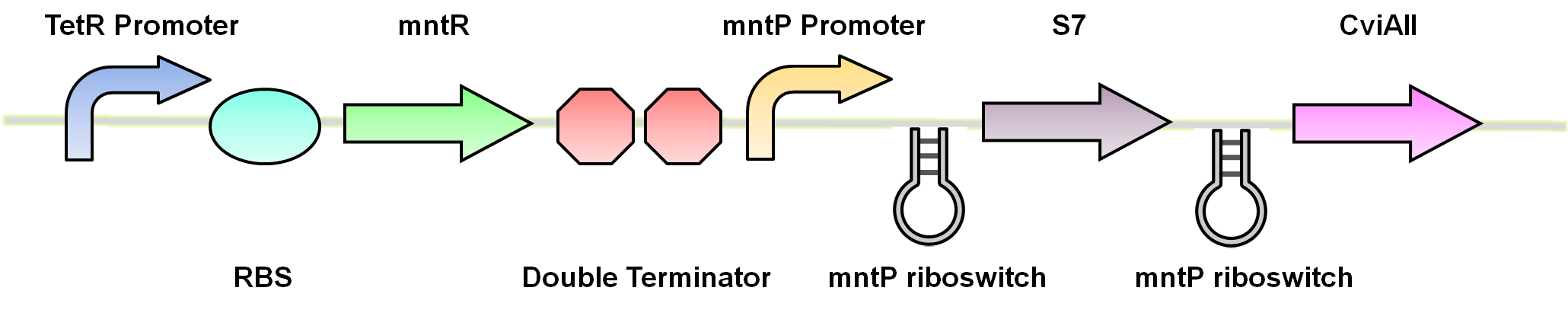
For our system we use the mntR,mntP promoter and mntP riboswitch. At high levels of manganese the mntP promoter mntPp and riboswitch mntPrb is activated producing the genes downstream. For our purposes we will be constructing mntPp and mntPrb with S7 and CviAII.
Test circuits for the manganese system
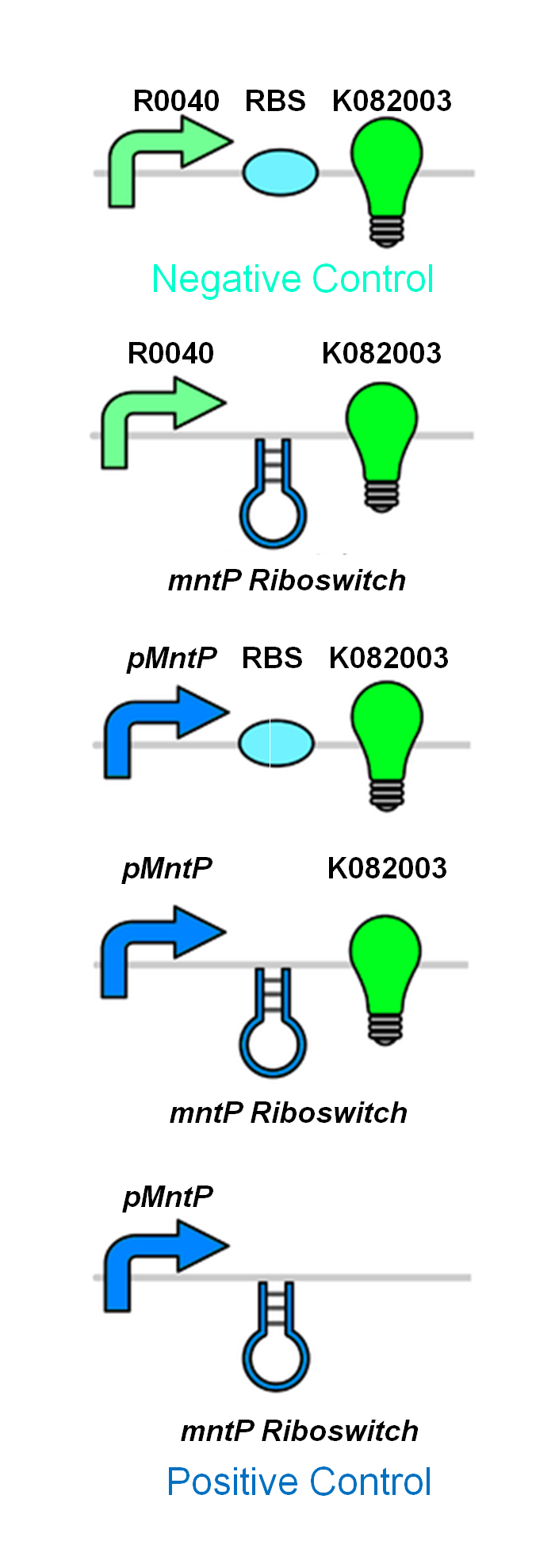
The manganese system was constructed with GFP-LVA for characterization purposes. The following are the control circuits built in order to characterise the MntP promoter and the MntP riboswitch.
Future Use
Even though this system is controlled and a good way to regulate kill genes, there are some limitations to this system. Tailings water contain high levels of Mn2+ in the contaminated water ~48mg/L or ~170µM/L. This concentration is 17 times the amount of Mn2+ needed to trigger the system (10 µM) therefore an additional chemical such as EDTA (a chelator) will have to be added to lower the manganese levels in the bioreactor. This however requires us to add EDTA into each set of tailings that is added into the bioreactor. EDTA is expensive and will thereby increase the cost of our bioreactor. Although this regulation method may not be feasible for our bioreactor, this regulatory system may be used in another pathway.
Glucose repressible system
Purpose
The rhamnose inducible promoter is suitable for controlling the kill switch in the the bioreactor since the promoter is shown to be tightly repressed with glucose. We aim to supplement the bioreactor with glucose to inhibit expression of the kill genes in the bioreactor. Escape of bacteria into the tailings ponds will cause expression of the kill genes since glucose levels in the surrounding environment would be insufficient for deactivate the promoter. We selected this system for two reasons. Firstly, the promoter exhibits tight repression with glucose and could thus limit inadvertent expression of the kill genes; and secondly, glucose is a common monosacharide which is economically feasible to continuously supplement into the bioreactor.
Application of the rhamnose promoter
The rhamnose promoter (pRha) is responsible for regulating six genes related to rhamnose metabolism and contains a separate promoter on its leading and reverse strands (see Figure one). RhaR and RhaS serve to regulate expression of the rhamnose metabolism genes rhaB, rhaA, and rhaD on the opposite side of the promoter. The RhaR transcription factor is activated by L-rhamnose to up-regulate expression rhaSR operon. In turn, the resulting RhaS activates the rhaBAD operon to generate the rhamnose metabolism genes (Egan & Schleif, 1993).
As a kill switch regulator, our team has harnessed global catabolite repression of the rhamnose promoter. Expression of the rhaBAD operon with RhaS requires the binding of catabolite receptor protein (CRP) cAMP complex to the promoter. When glucose is present, cAMP levels are low, such that CRP is unable to activate the promoter (Egan & Schleif, 1993). With this mechanism, our team set out to control our kill genes with glucose with the following rhamnose promoter construct:
Our kill system is different from the native rhamnose system with the rhaR and rhaS control genes. We have constituitively expressed rhaS to overcome dependency on rhamnose to cause activation of the kill switch. While RhaS is continuously present, global catabolite repression prevents its activation the kill genes in the bioreactor; in the outside environment, glucose levels are lower such that RhaS is able to activate the kill genes.
Components and characterization
Our team had pRha promoter commercially synthesized as per the sequence given by Jeske and Altenbuchner (2010). The rhaS and rhaR genes were amplified via PCR from Top Ten E. coli using Kapa Hi-Fi polymerase.
Given the control gene modifications which we have engineered into our system to optimize it for the tailings ponds, we are working to determine whether glucose repression of our modified system match patterns shown by Giacalone et al. (2006) and Jeske and Altenbuchner (2010). To this end, we have constructed the rhamnose promoter with GFP and are finalizing construction with constituitively expressed RhaS, so that we can characterize (LINK) this promoter and test it with our kill genes.
The Moco Riboswitch
Background
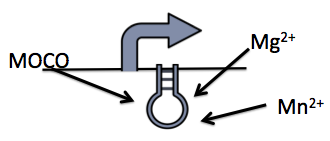
The molybdenum cofactor (moco) riboswitch is an RNA element which responds to the presence of the metabolite molybdenum cofactor (Regulski et al, 2008). This RNA element is located in the E.coli genome just upstream of the moaABCDE operon, which contain the important moco synthesis genes. Moco is an important cofactor in many different enzymes ranging from to this. The moco riboswitch has 2 regions: an aptamer domain and the expression platform. When moco is present in the cell it will bind to the aptamer region in the riboswitch which will cause an allosteric change. This allosteric change affects the expression platform by physically hiding the ribosome binding site thus preventing translation from occurring and hence adding a translational level of gene expression regulation. Therefore, the moco riboswitch, when activated by moco, inhibits gene expression.
The Killswitch Design
This riboswitch system can be used to design a killswitch mechanism to regulate the expression of our kill genes, CviAII and S7. The basic design of this system includes the kill genes downstream of the riboswitch all of which is under the control of a constitutive promoter. Downstream of this construct is the moa operon constitutively expressed. This entire construct will be present in a low copy plasmid in our bacteria. While moco synthesis is a normal process in the bacteria we wanted to constitutively express the moa operon to up regulate the expression of moco. This is to ensure high enough concentrations of moco present in order to inactivate the kill switch when the bacteria are in the bioreactor.
Molybdate in TPW
Molybdenum, Mo, is a trace element that is required by the bacteria for moco synthesis. Bacteria cannot uptake molybdenum in the elemental form and so uptakes molybdenum in its oxyanion form molybdate, MoO4, using the molybdate transport system. We wanted a system where molybdate is present in the bioreactor permitting moco synthesis (inactivate killswitch) and absent in the tailings pond water (TPW) preventing moco synthesis (activate killswitch). We discovered that molybdenum is indeed present in the TPW at around 1 µM.
Upon further literature research we found that molybdate is transported using a molybdate transport system which is coded by the modABCD operon. It has been shown that knocking out modC enables the transport system dysfunctional and prevents moco synthesis. However, if molybdate is supplemented in high enough concentrations to the bacteria, the bacteria are able to use its sulphate transport system to transport the molybdate in. This gave rise to the idea of having a modC knock out strain of bacteria in our system and supplementing it with molybdate. The paper tried 10mM sodium molybdate supplementation in the media to repress gene expression whereas that the literature level states that there is less than 1 µM molybdate in the TPW. Therefore, in the tailings ponds, the concentration of molybdate is not high enough to be transported into the cell using the sulphate transport mechanism. This prevents moco synthesis and activates the killswitch. While sodium molybdate is expensive and can be argued to be impractical to provide on a large bioreactor scale, there are many inexpensive and effective filtration mechanisms available to filter out most of the molybdate and reuse it.
Characterization
There are two experiments that we want to run with this system. First, we wish to characterize the concentration of molybdate needed in the system to enter through the sulphate transport mechanism. We will supply lab strain E.Coli and modC knockout strain of E.Coli with varying concentrations of molybdate and measure the optical density. This will be followed by a CFU assay. This experiment will also inform us of toxic levels of molybdate for our bacteria. Our second experiment is the characterization of the moco killswitch system.
Final Moco Construct:

 "
"