Team:Calgary/Notebook/Killswitch
From 2012.igem.org
| Line 155: | Line 155: | ||
<h2>Week 10 (July 3-July 6)</h2> | <h2>Week 10 (July 3-July 6)</h2> | ||
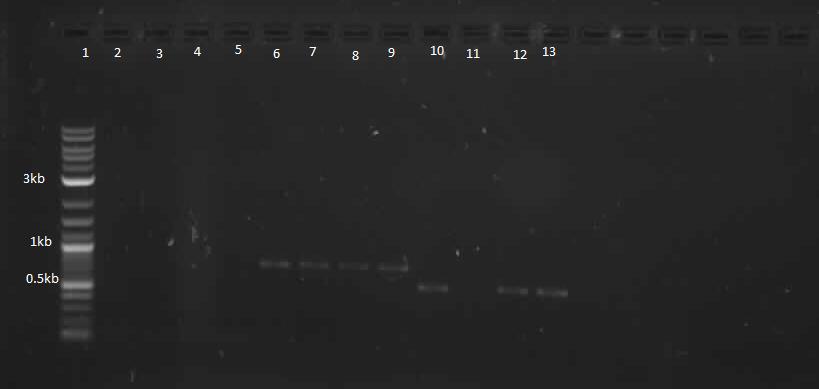
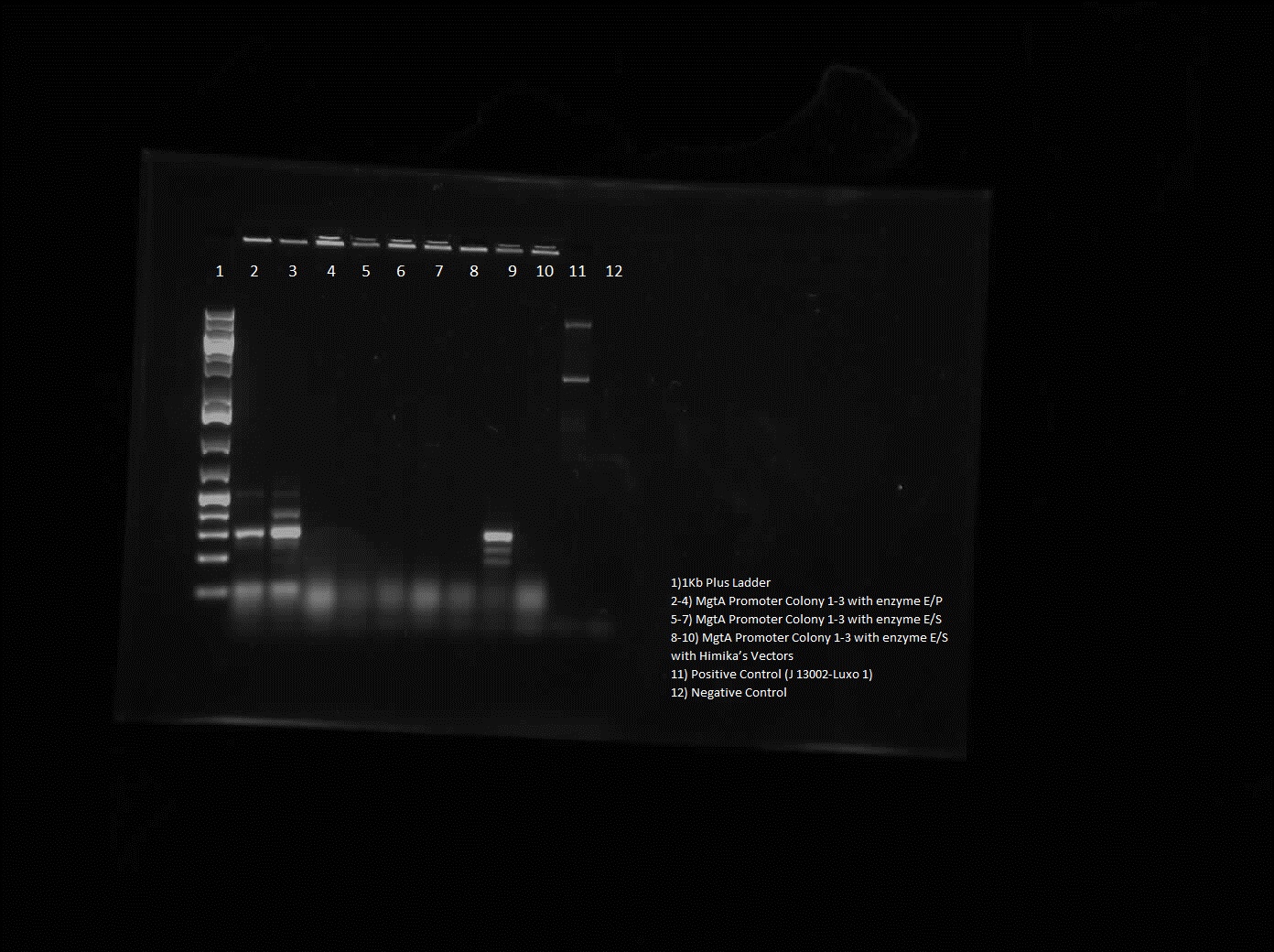
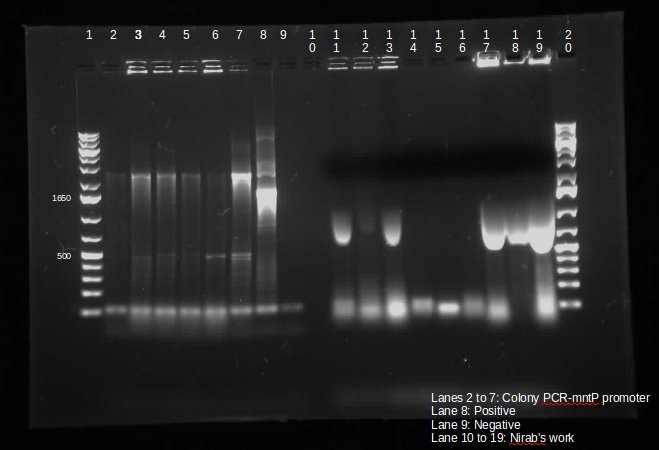
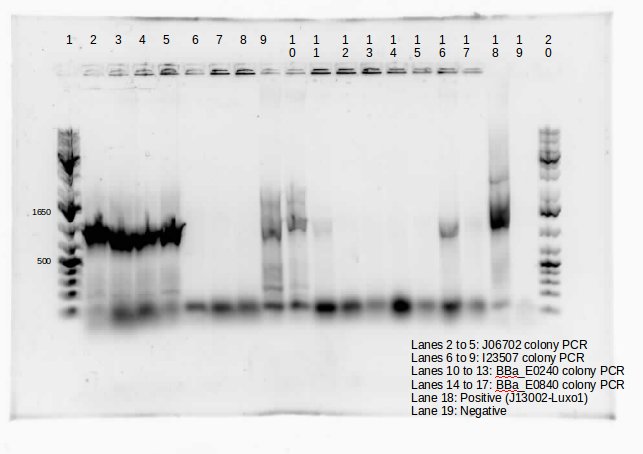
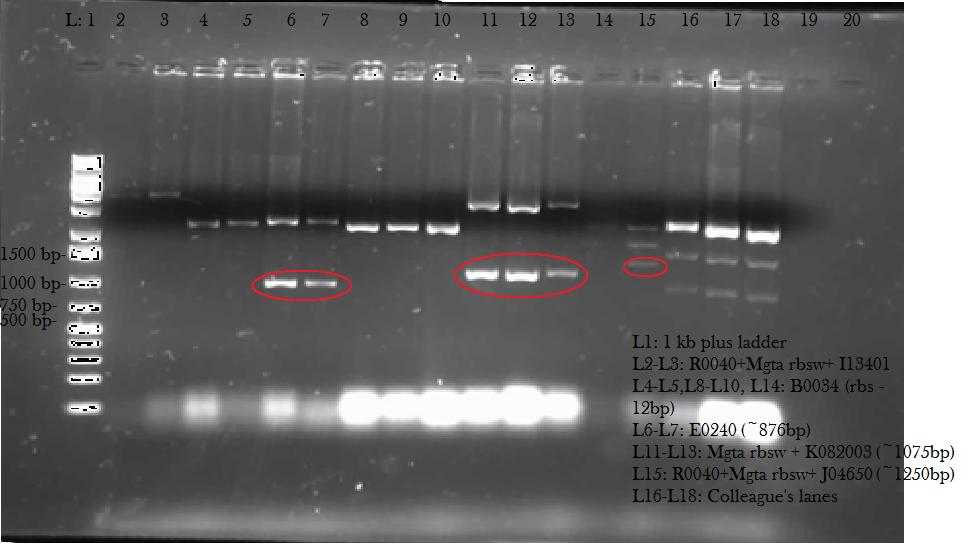
| - | <p> This week, we attempted to religate our constructions from the previous week as they were not verified as positive in our colony PCR. We had run a gel to see if the inserts and vectors were digested properly and they were. Following transformation, we verified it through cPCR. The two gels show the results of construction involving two promoters (Mgta Promoter and a constitutive tetR promoter, R0040) and the mgta rbsw with different reporter gene constructs from the previous week. There are also some plasmid switches and constructions of the promoters and just the riboswitch. Based on these results, we isolated plasmids of R0040 + (Mgta RBSW and I13401) from L6 and L7 as well as I13507 from L8 from the first gel and R0040 + (Mgta RBSW and J04650) from the second gel (circled in red). We verified these constructions with a digest. The third gel shows the digest and the lanes circled are the ones that look good and were sent to sequencing. </p> | + | <p> This week, we attempted to religate our constructions from the previous week as they were not verified as positive in our colony PCR. We had run a gel to see if the inserts and vectors were digested properly and they were. Following transformation, we verified it through cPCR. The two gels show the results of construction involving two promoters (Mgta Promoter and a constitutive tetR promoter, R0040) and the mgta rbsw with different reporter gene constructs from the previous week. There are also some plasmid switches and constructions of the promoters and just the riboswitch. Based on these results, we isolated plasmids of R0040 + (Mgta RBSW and I13401) from L6 and L7 as well as I13507 from L8 from the first gel and R0040 + (Mgta RBSW and J04650) from the second gel (circled in red). </p> |
| + | |||
| + | </html>[[File:iGEMCalgary2012.07.04.2012 Construction cPCR.jpg|thumb|400px|center|Gel 1: Colony PCR of Constructions]]<html> | ||
| + | </html>[[File:iGEMCalgary2012.07.04.2012 Construction cPCR 2 (1) with label.jpg|thumb|400px|center|Gel 2: Colony PCR of Constructions]]<html> | ||
| + | |||
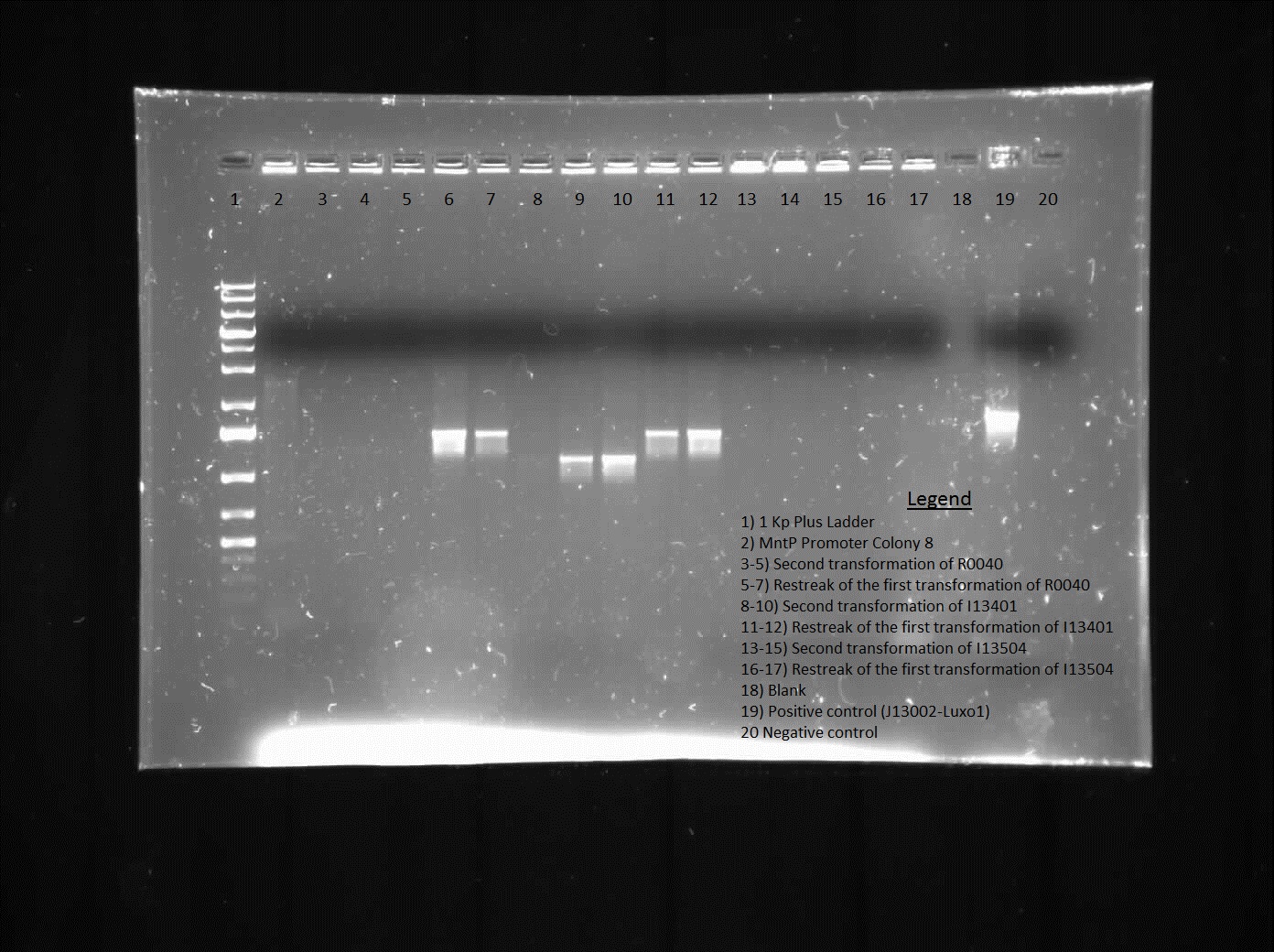
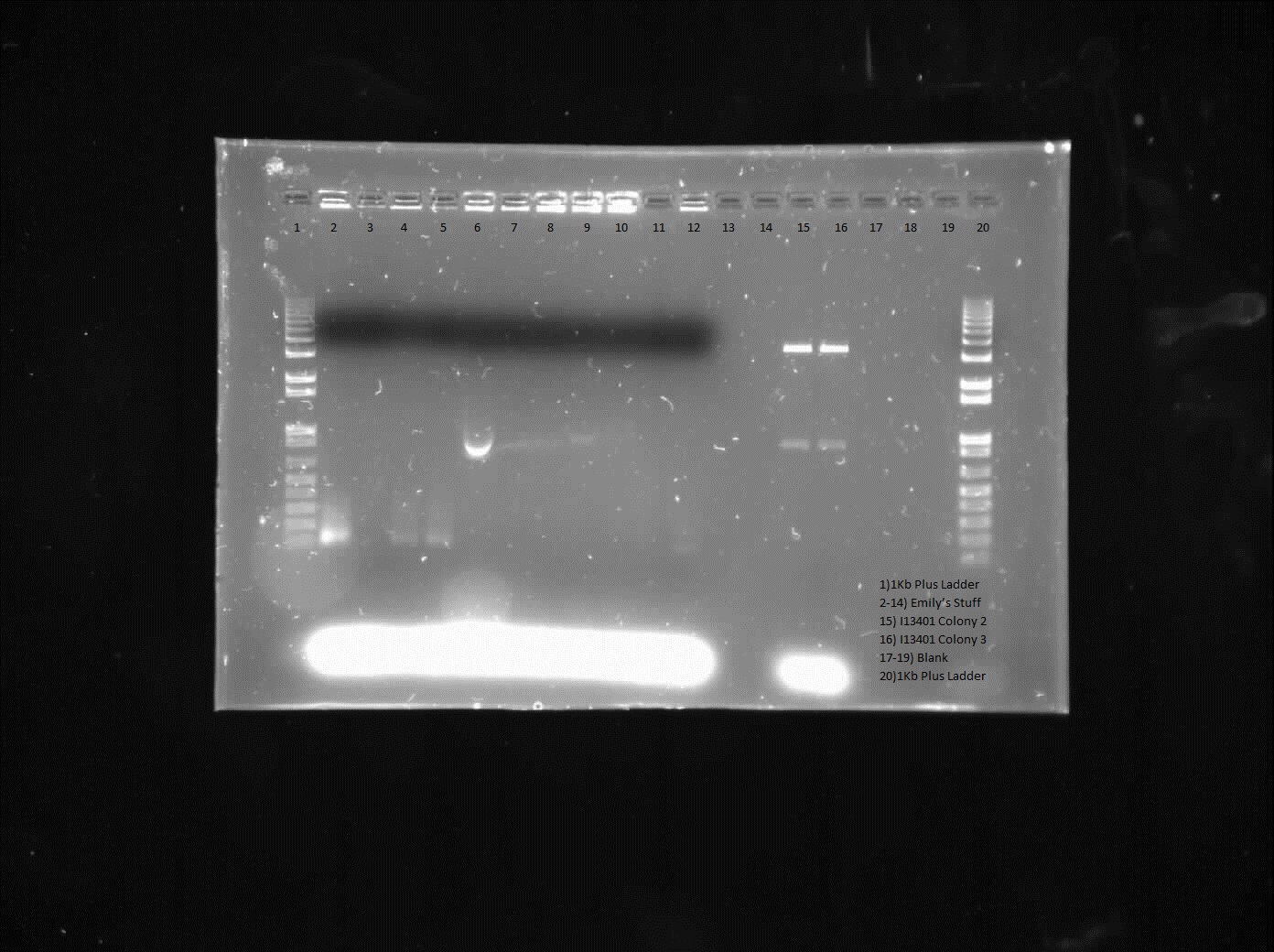
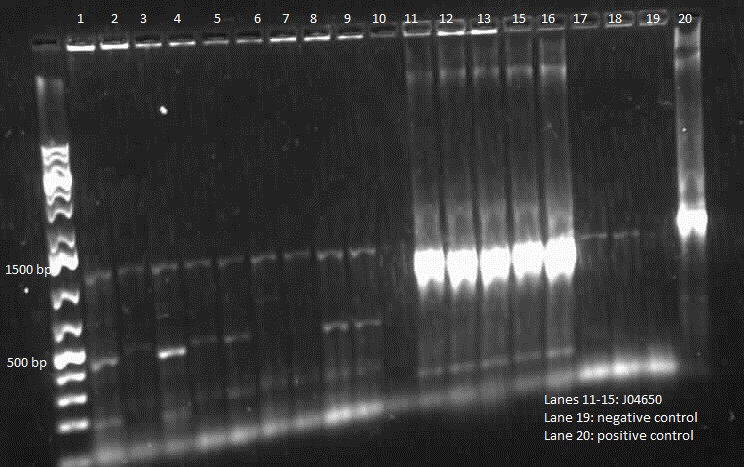
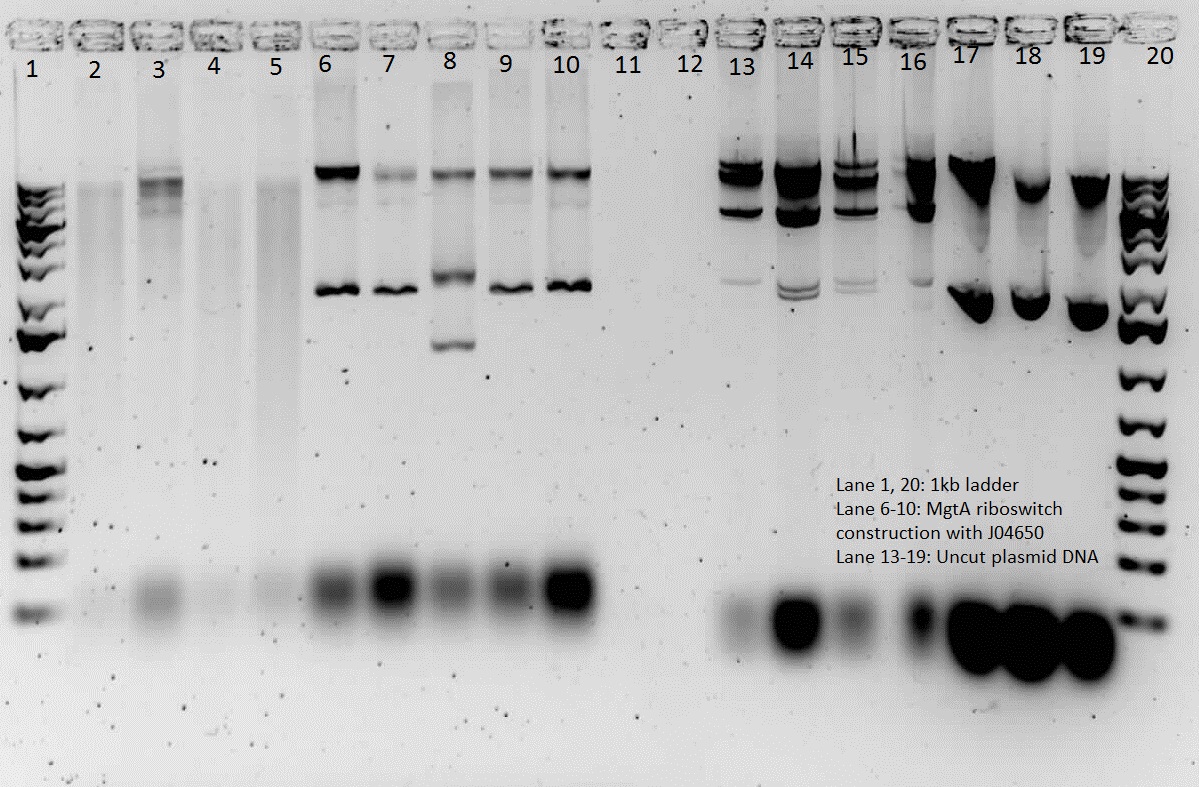
| + | <p> We verified these constructions with a digest. The third gel shows the digest and the lanes circled are the ones that look good and were sent to sequencing. </p> | ||
| + | |||
| + | </html>[[File:iGEMCalgary2012.07.06.2012 verification of overnight digest with labels.jpg|thumb|400px|center|Verification digest of constructions]]<html> | ||
Revision as of 17:52, 23 July 2012
Week 1 (May 1-4)
This was the first week where we met with other team members and summarized the primary subprojects the team will be tackling this coming summer.
Week 2 (May 7-11)
Members were assigned to the killswitch team. We spent a majority of this week performing literature searches and narrowing the killswitch to a few ideas.
The mechanism of death is still settled upon the micrococcal nuclease, but regulation of the death genes will be difficult. Existing repressible promoters in the registry still tend to be leaky in their expression, but we need to test this out. We may look into the TetR-repressible promoter (R0040) and also the clλ regulated promoter (R0051).
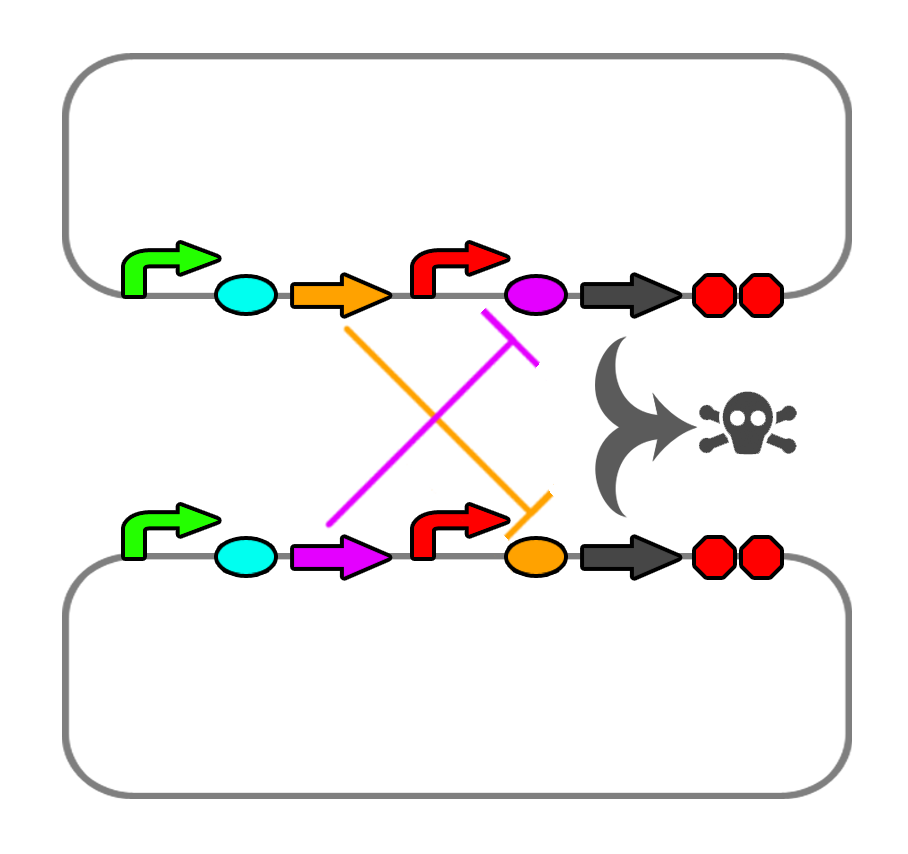
We also found a few riboswitches responsible to various metal ions such as Mg2+ and Mn2+. The mgtA riboswitch activates translation of the death gene when magnesium ions are not present in the solution. The mntA riboswitch deactivates translation of the death gene when manganese ions are not present. Considering this, we may be able to find a method of precipitating or otherwise sequestering Mn2+ ions out of the tailings water prior to entering the bioreactor. This way, our bacteria would not die when they come into contact with the manganese in the tailings water. Possible additives include carbonate (CO32-) or hydroxide (OH-), though this may alter the pH too greatly.
One other proposal would be to use promoters that activate when the bacteria detect the formation of a biofilm on, for example, glass beads. Engineering a monolayer of cells on a bead means that if a cell detaches, its death genes will activate. Further research into this is needed.
Week 3 (May 14-18)
We are continuing to look in to various other pathways. We discussed the possibility of a NOR-gated system to add a second layer of regulation to the kill gene. It would require the production of riboswitch ligands. Possible ligands include glucose, amino acids, and molybdenum cofactor.
We also looked into a co-dependent system where we would transform E. coli with two different plasmids and put them in the same bioreactor such that they produce ligands for each other that would repress the expression of the restriction enzyme and the nuclease. For this we explored riboswitches further and came up with the idea of using MOCO and SAM riboswitches as SAM is soluble in water and not found in the tailings ponds.
We also found a glucose repressible promoter that we could have potentially used upstream of the kill switch. This promoter is found in F. tularensis. However we also found that this promoter is not found in E. coli and F. tularensis has a unique polymerase which used this promoter. Hence using this promoter does not seem feasible. Next week we will be looking into further possibilities for the kill switch.
Week 4 (May 21-25)
In terms of our literature search we decided against the glucose repressible promoter from F. tularensis because it was not native to E.coli and upon contact with the authors of the paper we found that the promoter did not work in E.coli. We also came upon two systems that we are interested in further investigating. The first one was a glucose activated promoter that was found in the registry which we thought of coupling to an inverter and the second one is a rhamnose-based regulatory system.
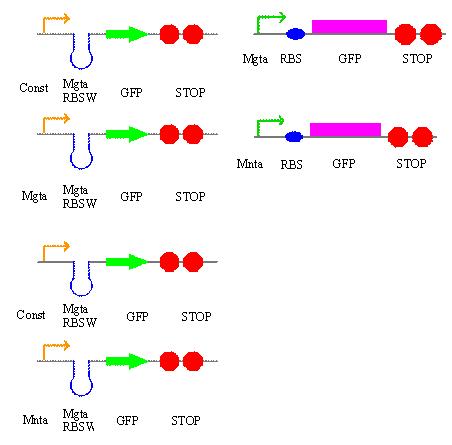
On Tuesday, in addition to our literature research, we decided on some of the circuits that we will be building over the next couple of weeks and came up with a plan detailing what will be done each day. These are the circuits:
On Wednesday we proceeded with the transformation of 3 parts from the iGEM kit into our E.coli: a tetR promoter, a GFP-double terminator, and a RBS-GFP-double terminator. On Thursday we verified our transformation process with a cPCR where we ran 10 colonies from each plate with our controls. We ran a gel on the PCR products and viewed it to verify the transformation and to decide which of the successful colonies to plate and grow overnight.
On Friday we isolated the plasmids through a mini prep. The plan for next week is to do a restriction digest and run it on a gel to confirm that the plasmids have the part. We also want to start more transformation of our parts into E.coli such as the riboswitches and the promoters. We want to verify the transformation, isolate the plasmids and verify, and start the DNA construction digest to start building our circuit.
Week 5 (May 28-June 1)
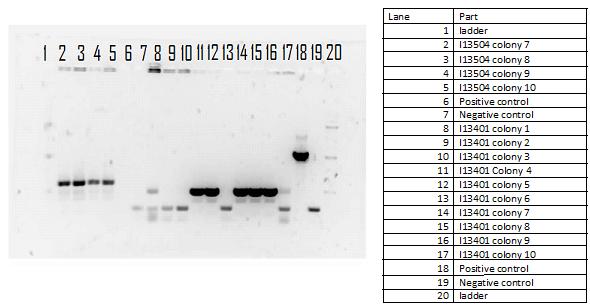
During the week of May28th, the killswitch team made progress toward transforming/verifying nine different parts in E. coli. Work began with continuation of the previous week's transformation of R0040 (TetR promoter), I13401 (GFP terminator), and I13504 (RBS GFP terminator). Note that these three parts will be necessary for characterization of the four different kill system triggers being explored by the killswitch team. Verification procedures (ie, colony PCR) showed successful transformation of I13401. Results for the other two parts were inconclusive and cloning of R0040 and I13504 will continue next week.
GEL SHOWING SUCCESS OF I13401
The commercially synthesized mntA promoter, mgtA promoter, and mgtA riboswitch for the magnesium and manganese riboswitches were also transformed into E. coli. While colonies were successfully isolated, colony PCR was inconclusive because incorrect primers were used for the commercial plasmids. The team proceeded to isolate the plasmids from E. coli, perform a restriction digest of the parts, and ligation of these components into biobrick vectors. These products were transformed into other E. coli and the team is ready to verify presence of the cloned digest products.
In continuation with magnesium and manganese riboswitches, the team attempted to biobrick mntR (a repressor of a manganese transporter activated by manganese), PhoP (a transcriptional regulator of mgtA), and PhoQ (a signal transducer which activated PhoP in the presence of divalent cations). The below PCR indicates that while PhoQ and mntR were successfully isolated, PhoP was not. The former two were transformed into E. coli; the transformed samples grew into a lawn and colony PCR will thus be performed next with re-streaked plates. Finally, gradient PCR was used in an attempt to biobrick PhoP.
Lanes 6-9: PhoQ (~671 bp)
Lanes 10-13: mntR (~500 bp)
Primers for the MOCO and rhamnose killswitch actuators were also designed. With respect to the MOCO synthesis operon and MOCO riboswitch, primers were finalized and sent to senior members of the team. While primers for isolation of rhaSR and the rhaT promoter from E. coli were given in the paper which inspired the rhamnose promoter idea, the team was resistant to using the sequences since they did not seem to correlate with genes of interest in a BLAST search. Closer inspection revealed that the forward primer listed in the paper added a number of bases (>50) which were not native to the E. coli genome. Only 18 base pairs of the primer were complementary to E. coli—this discrepancy was responsible for the negative results of the BLAST alignment. Work will continue next week to optimize the rhamnose primers. They hope to order both the MOCO and rhamnose primers next week so that they may proceed with construction of the kill switch.
Week 6 (June 4th-8th)
This week we continued our attempt to biobrick PhoP, PhoQ and mntR. A gradient PCR was run with temperature and salt gradient (0.5mM-3.0mM). however this PCR showed no success whatsoever. We discovered that the primers designed for these parts were faulty and were binding in several places in the gene and thereby favouring products that are smaller in size.
phoP-S GTTTCTTCGAATTCGCGGCCGCTTCTAGatgCGCGTACTGGTTGTTG
phoP-AS GTTTCTTCCTGCAGCGGCCGCTACTAGTATTATTAtcaGCGCAATTCGAACAGATAG
phoQ-S GTTTCTTCGAATTCGCGGCCGCTTCTAGatgaaaaaattactgcgtc
phoQ-AS GTTTCTTCCTGCAGCGGCCGCTACTAGTATTATTAttattcatctttcggcgcag
mntR-S GTTTCTTCGAATTCGCGGCCGCTTCTAGatgagtcgtcgcgcaggtac
mntR-AS GTTTCTTCCTGCAGCGGCCGCTACTAGTATTATTAtcatttggcaccgtgtttc
Hence new primers were designed.
PhoPNEW-AS CTGCAGCGGCCGCTACTAGTAGCAGTAATTTTTTCATCAGCGCAATTCG
PhoQNEW-AS CTGCAGCGGCCGCTACTAGTAGCATATTTATTCATCTTTCGG
mntRNEW-S gaattcgcggccgcttctagGAGGAAGCACAATGAGTC
mntRNEW-AS CTGCAGCGGCCGCTACTAGTAGCGTGCGTAAAAAAGG
The killswitch team also focused on verification of both the magnesium/manganese (MgtA promoter, MgtA riboswitch and MntP promoter) and registry parts (R0040, I13401 and I13504).
For the magnesium/manganese parts this week we insert them in biobrick vectors in order to carry out colony PRR. Colony PCR of the magnesium/manganese parts could not have been done first since we did not have the right primer , therefore by doing a construction digest we could use the standard biobrick primers. From the gel, the results were inconclusive as the ladder was not clear and the parts were too small to tell the difference between them and primer dimers, therefore next week goal is to perform another CPCR of the parts. As well as doing colony PCR of the MgtA promoter, a restriction digest of this part was done. The gel showed good results as the bands were bright and the right size.
For the registry parts R0040 no results were shown in the initial colony PCR, however another gel was done with R0040 from the 2010 and 2011 registry plate. The colony PCR of R0040 from 2011 showed the rights bands, however the results are inconclusive as the negative was contaminated, therefore most likely another colony PCR needs to be done.
The registry part I13401 had a successful week as both the colony PCR and the restriction digest verified our parts. Since the result was successful we performed glycerol stocks of the plasmid and prepared it for sequencing the following week.
The part I13504 showed no results this week therefore the goal next week is to find another part that is similar to I13504 and test its verification.
Week 7 (June 11-14)
This week we continued the insertion of magnesium promoter (MgtA) into biobrick plasmid (PSB1C3) to carry out the colony PCR to confirm the presence of the Mgta part in the plasmid using Bbk_CP_F and Bbk_CP_R primers. We are currently in the process of further verification by using a mini prep. We also transformed four registry parts (J06702 [RBS-mCherry-dTerm], I13507 [RBS-RFP-dTerm], E0840 [RBS-GFP-dTerm] and E0240 [RBS-GFP-dTerm]) directly from the kit plates and verified the parts using colony PCR the following day. The primers used to verify these parts were Bbk_CP_F and Bbk_CP_R. These parts were successfully transformed but we still saw some contaminant bands in the gels (see below).
]
Transformation of MntP in PSB1C3 vector was verified using colony PCR. However, the results indicated that the presence of MntP as well as a contaminant band around 3000 bp.
]Transformation of registry parts I13521 [TetR-B0034-RFP-dTerm], I13507 and J04650 [RFP-dTerm] were performed and colony PCR was carried out to verify the presence of the parts in the cells. J04650 was the only part that was successfully transformed. Though the other parts were not successfully transformed, we can still see non-specific bands in the gel (see below). We are currently doing mini prep to additionally verify the success of transformation.
Week 8 (June 18-June 22)
This week we did a restriction digest of all the plasmids that were isolated. We confirmed that we had J04650, I13507, E0240 and mgtA riboswitch. Hence these were sent to sequencing. We got sequencing results back and confirmed that we have J04650, I13507, J06702, R0040, E0240 and mgtA riboswitch. Henceforth, we started constructing the mgtA riboswitch with J04650.
The mgtA riboswitch with J04650 was confirmed using PCR as well as restriction digest. We plan on sequencing it when we are able to construct it with a promoter.
Week 9 (June 25-June 29)
This week we attempted to construct the mgtA riboswith and J04650 construct with promoters such as MgtA and TetR. We also tried to plasmid switch I13507 such that we can put mgtA promoter on to that composite part. So far we have verified through colony PCR that the
Week 10 (July 3-July 6)
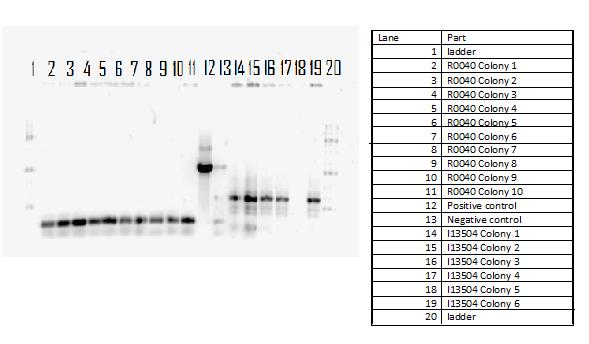
This week, we attempted to religate our constructions from the previous week as they were not verified as positive in our colony PCR. We had run a gel to see if the inserts and vectors were digested properly and they were. Following transformation, we verified it through cPCR. The two gels show the results of construction involving two promoters (Mgta Promoter and a constitutive tetR promoter, R0040) and the mgta rbsw with different reporter gene constructs from the previous week. There are also some plasmid switches and constructions of the promoters and just the riboswitch. Based on these results, we isolated plasmids of R0040 + (Mgta RBSW and I13401) from L6 and L7 as well as I13507 from L8 from the first gel and R0040 + (Mgta RBSW and J04650) from the second gel (circled in red).
We verified these constructions with a digest. The third gel shows the digest and the lanes circled are the ones that look good and were sent to sequencing.
 "
"