Team:Bielefeld-Germany/Test
From 2012.igem.org
(Difference between revisions)
| Line 1: | Line 1: | ||
| - | + | ==Week 25 (10/15 - 10/21/12)== | |
| + | |||
| - | |||
| + | __NOTOC__ | ||
| + | <html> | ||
| + | <table> | ||
| + | </table> | ||
| + | </html> | ||
| - | ===Monday October | + | ===Monday October 15th=== |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
* '''Team Shuttle Vector:''' | * '''Team Shuttle Vector:''' | ||
| - | ** | + | ** There are no colonies of pBS1C3::BBa_K863202 KRX transformantion. |
| + | ** The P. pastoris GS115 cells integrated with BBa_K863204::GFP were harvested and the supernatant was examined on GFP by fluorescence. But there was no GFP. The next steps are: disrupt the cells and test the supernatant on GFP. Maybe the GFP could not be secreted. | ||
| + | * '''Team Fungal and Plant Laccases:''' | ||
| + | ** Yeah, the yeast cells are growing. | ||
* '''Team Cellulose Binding Domain:''' | * '''Team Cellulose Binding Domain:''' | ||
| - | ** | + | ** Tried to isolate the eight plated, but did get low concentrations again (except one J61101+GFP_Freiburg+CBDcex_Freiburg with conc >100 ng/µL) |
| - | ** | + | ** cleaned up some more PCR-products (GFP_Freiburg; CBDcex_Freiburg; CBDclos_Freiburg) |
| - | + | *'''Team Substrate Analysis''': We stopped the reaction from the 12th October with Methanol and gave it to Dr. Marcus Persicke to measure the degradation on LC-MS | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | * '''Team | + | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | === | + | ===Tuesday October 16th=== |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | * | + | * '''Team Activity Tests:''' |
| + | ** Today we received the samples from Team Cultivation & Purification. We are really motivated to measure the activity of our new produced enzymes. But before that we have to prepare the samples. This implied the determination of the protein concentration in each fraction, which we did today. The results are listed in the following table: | ||
| + | [[File:Bielefeld2012_Prot_vor_Umpuffern.jpg|300px|thumb|center]] | ||
| - | |||
* '''Team Cellulose Binding Domain:''' | * '''Team Cellulose Binding Domain:''' | ||
| - | ** | + | ** Transformed B0034 into KRX |
| - | ** | + | ** Transformed J23100 into KRX |
| - | + | ** Plated both on AMP-select-agar | |
| - | ** | + | ** Plated one more Colony of J61101 + GFP_Freiburg and J61101 + GFP_Freiburg + CBDclos |
| - | + | region=DE Laccase] || 150 µL || | |
| - | + | |- | |
| - | + | |} | |
| - | ** | + | *'''Team Substrate Analysis:''' The Massspectrometry data showed two potential products after the Laccase treatment of Estradiol and Ethinyl estradiol but we unfortunatly could not identify them so we decided to do MS-MS on thoose. The products showed that after Laccase treatment both Estradiol and Ethinyl estradiol losses two '''H''' atoms which was new for us. |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | * '''Team | + | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | * '''Team | + | ===Wednesday October 17th=== |
| - | ** | + | * '''Team Activity Tests:''' |
| - | + | ** Before starting our activity assay we re-buffered the fractions we got from Team Cultivation and Purification. It took some time, that's why we had no time for activity analysis yet. But we prepared everything for the next day, including determining the protein concentration of the fractions after they have been re-buffered. This is what we got: | |
| - | + | [[File:Bielefeld2012_Prot_nach_Umpuffern.jpg|300px|thumb|center]] | |
| - | + | ||
| - | + | ||
| - | + | ||
* '''Team Shuttle Vector:''' | * '''Team Shuttle Vector:''' | ||
| - | ** | + | ** Genomic DNA isolation of yeast cells with integrated BBa_K863204::GFP construct was done with the Kit. For genotype characterisation an PCR with the praimer pair 5AOX-Genotyp-FW and TT-Genotyp-RV was done and the fragment size was determined. |
| + | * '''Team Fungal and Plant Laccases:''' | ||
| + | ** YPD liquid culture was inoculated with one yeast colony (BBa_K863204::TV5) and incubated at 30°C and 150 rpm. | ||
| + | * '''Team Cellulose Binding Domain:''' | ||
| + | ** Picked colonies of B0034 and J23100 and plated them on AMP selection agar for plasmid isolation | ||
| + | * '''Team Site Directed Mutagenesis:''' | ||
| + | ** SDM-PCR of the Shuttle vector | ||
| + | ** Added ''Dpn''I over night | ||
| - | * '''Team | + | ===Thursday October 18th=== |
| - | ** | + | * '''Team Activity Tests:''' |
| + | ** Finally the day had come to measure the activity of our own produced laccases. This activity assay should give information about the enzyme content in every fraction. To make the measurements comparable we applied the same protein amount in every sample. Again, we used our [https://2012.igem.org/Team:Bielefeld-Germany/Protocols/Analytics#General_setup_of_enzyme_activity_measurements/ standard activity assay protocol], but this time we used the [https://2012.igem.org/Team:Bielefeld-Germany/Protocols/Materials#Britton-Robinson_Buffer/ Britton-Robinson Buffer] at pH 5. Also we applied 0.1 mM ABTS and measured over night. | ||
| + | [[File:Bielefeld2012_new_ECOL_activity.jpg|360px|thumb|left|Activity assay of each purified fraction of our new cultivation with ECOL. Samples were re-buffered into H<sub>2</sub> and the protein amount in each fraction has been adjusted. The Measurement was done using the [https://2012.igem.org/Team:Bielefeld-Germany/Protocols/Analytics#General_setup_of_enzyme_activity_measurements/ standard activity assay protocol] over night.]][[File:Bielefeld2012_new_BPUL_acitivity.jpg|360px|thumb|right|Activity assay of each purified fraction of our new cultivation with BPUL. Samples were re-buffered into H<sub>2</sub> and the protein amount in each fraction has been adjusted. The Measurement was done using the [https://2012.igem.org/Team:Bielefeld-Germany/Protocols/Analytics#General_setup_of_enzyme_activity_measurements/ standard activity assay protocol] over night.]] | ||
| + | [[File:Bielefeld2012_new_BHAL_activity.jpg|360px|thumb|left|Activity assay of each purified fraction of our new cultivation with BHAL. Samples were re-buffered into H<sub>2</sub> and the protein amount in each fraction has been adjusted. The Measurement was done using the [https://2012.igem.org/Team:Bielefeld-Germany/Protocols/Analytics#General_setup_of_enzyme_activity_measurements/ standard activity assay protocol] over night.]][[File:Bielefeld2012_new_TTHL_activity.jpg|360px|thumb|right|Activity assay of each purified fraction of our new cultivation with TTHL. Samples were re-buffered into H<sub>2</sub> and the protein amount in each fraction has been adjusted. The Measurement was done using the [https://2012.igem.org/Team:Bielefeld-Germany/Protocols/Analytics#General_setup_of_enzyme_activity_measurements/ standard activity assay protocol] over night.]] | ||
| + | <br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
* '''Team Cellulose Binding Domain:''' | * '''Team Cellulose Binding Domain:''' | ||
| - | ** | + | ** Isolated J23100 plasmid |
| - | ** | + | ** Digested J23100 with ''Eco''RI and ''Pst''I |
| + | ** Digested pSB1C3-Backbone with ''Eco''RI and ''Pst''I | ||
| + | ** Clean up of pSB1C3-BB and J23100 + RFP - Insert via Gel | ||
| + | ** Ligation of J23100 and pSB1C3-BB | ||
| + | ** Isolated B0034 plasmid | ||
| + | ** Digested B0034 with ''Spe''I and ''Pst''I | ||
| + | ** Digested GFP_Freiburg-PCR-product with ''Xba''I and ''Pst''I | ||
| + | ** Digested an other fraction of the GFP_Freiburg-PCR-product with ''Xba''I and ''Age''I | ||
| + | ** Assembled (Ligation): | ||
| + | *** B0034 with GFP_Freiburg | ||
| + | *** B0034 with GFP_Freiburg and CBDcex_Freiburg | ||
| + | *** B0034 with GFP_Freiburg and CBDclos_Freiburg | ||
| + | ** Transformed these three into KRX | ||
| - | * '''Team | + | * '''Team Site Directed Mutagenesis:''' |
| - | ** | + | ** Bands of 19C-PRC-product in Gel at 7 kbp (which is correct) |
| + | ** Clean-Up of 19C-PRC-product | ||
| + | ** Transformation of the 19C-PRC-product in XL1 Blue | ||
| + | ===Friday October 19th=== | ||
| + | * '''Team Activity Tests:''' | ||
| + | ** After getting our interesting activity measurement results, we had to figure out which fraction is going to be used in the following experiments and how much laccase it contains besides other proteins. We agreed, that the most active fraction contains 90 % laccase, which is commonly used in the literature. Since we applied the same protein amount of each fraction for the activity measurements, we made sure that the results really correspond to the amount of laccase. These are the most active fractions we have chosen with their contained laccase amount: | ||
| + | *** ECOL: Fraction 50% 2 with a laccase concentration of 63,9 µg mL<sup>-1</sup>. | ||
| + | *** BPUL: Fraction 50% 2 with a laccase concentration of 25,1 µg mL<sup>-1</sup>. | ||
| + | *** BHAL: Fraction 5% 3 with a laccase concentration of 10,9 µg mL<sup>-1</sup>. | ||
| + | *** TTHL: The best fraction of our TTHL laccase was fraction 50% 1 with a laccase concentration of 4,03 µg mL<sup>-1</sup>. In regard to Team Immobilization and Team Substrate Analysis we had to reach a higher amount of TTHL laccase. So we thought of combining two fractions. We chose the second best fraction of TTHL, which is 5% 3. To calculate the amount of contained laccase we had to compare the activity and reconsidered the slope. Since we stated the fraction with its highest activity as 90 % contained laccase, we calculated the amount of laccase regarding to that one. The following figure shows the slopes we got out of the activity from fraction 50% 1 and fraction 5% 3 (which is the second best). Accordingly to this, fraction 5% 3 contains 68,9% of laccase, which is 4,8 µg mL<sup>-1</sup>. In total, the composed fraction contains 4,4 µg mL<sup>-1</sup>. | ||
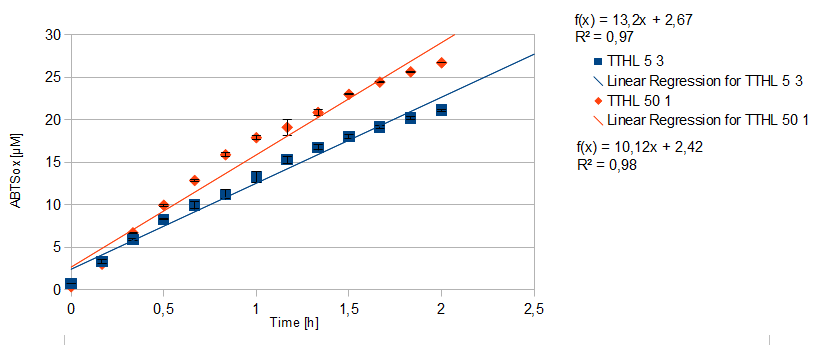
| + | [[File:Bielefeld2012_Regression.png|center|450px|thumb|Slope of the two most active fractions of purified TTHL laccase. By comparing them, the amount of contained laccase in the fractions had been calculated.]] | ||
| + | * '''Team Cellulose Binding Domain:''' | ||
| + | ** Colony-PCR of B0034 + GFP_Freiburg; B0034 + GFP_Freiburg + CBDcex; B0034 + GFP_Freiburg + CBDcex_Freiburg; B0034 + GFP_Freiburg + CBDclos_Freiburg. | ||
| + | *** All negative | ||
| + | ** Restriction of Ecol_Freiburg with ''Xba''I, ''Age''I and ''Dpn''I | ||
| + | * '''Team Site Directed Mutagenesis:''' | ||
| - | + | ===Saturday October 20th=== | |
| - | + | ===Sunday October 21st=== | |
| - | + | * '''Team Cellulose Binding Domain:''' | |
| + | ** Colonies PCRs of B0034 + GFP_Freiburg; B0034 + GFP_Freiburg + CBDcex_Freiburg; B0034 + GFP_Freiburg + CBDclos_Freiburg | ||
| + | *** All negativ | ||
| + | ** Transformed once again (with an mix of old an new GFP_Freiburg) | ||
| + | ** Isolated plated colonies of transformed colonies (two GFP; two GFP+Cex; two GFP+Clos | ||
| + | ** Restriction analysis showed only bands at 2K bp (pSB1C3+BBa_B0034 vector without insert) | ||
| + | * '''Team Site Directed Mutagenesis:''' | ||
| + | ** Isolated the plasmids of the six SDM-dishes | ||
| + | ** Over-night digestion with ''Not''I | ||
Revision as of 09:56, 26 October 2012
Week 25 (10/15 - 10/21/12)
Monday October 15th
- Team Shuttle Vector:
- There are no colonies of pBS1C3::BBa_K863202 KRX transformantion.
- The P. pastoris GS115 cells integrated with BBa_K863204::GFP were harvested and the supernatant was examined on GFP by fluorescence. But there was no GFP. The next steps are: disrupt the cells and test the supernatant on GFP. Maybe the GFP could not be secreted.
- Team Fungal and Plant Laccases:
- Yeah, the yeast cells are growing.
- Team Cellulose Binding Domain:
- Tried to isolate the eight plated, but did get low concentrations again (except one J61101+GFP_Freiburg+CBDcex_Freiburg with conc >100 ng/µL)
- cleaned up some more PCR-products (GFP_Freiburg; CBDcex_Freiburg; CBDclos_Freiburg)
- Team Substrate Analysis: We stopped the reaction from the 12th October with Methanol and gave it to Dr. Marcus Persicke to measure the degradation on LC-MS
Tuesday October 16th
- Team Activity Tests:
- Today we received the samples from Team Cultivation & Purification. We are really motivated to measure the activity of our new produced enzymes. But before that we have to prepare the samples. This implied the determination of the protein concentration in each fraction, which we did today. The results are listed in the following table:
- Team Cellulose Binding Domain:
- Transformed B0034 into KRX
- Transformed J23100 into KRX
- Plated both on AMP-select-agar
- Plated one more Colony of J61101 + GFP_Freiburg and J61101 + GFP_Freiburg + CBDclos
region=DE Laccase] || 150 µL || |- |}
- Team Substrate Analysis: The Massspectrometry data showed two potential products after the Laccase treatment of Estradiol and Ethinyl estradiol but we unfortunatly could not identify them so we decided to do MS-MS on thoose. The products showed that after Laccase treatment both Estradiol and Ethinyl estradiol losses two H atoms which was new for us.
Wednesday October 17th
- Team Activity Tests:
- Before starting our activity assay we re-buffered the fractions we got from Team Cultivation and Purification. It took some time, that's why we had no time for activity analysis yet. But we prepared everything for the next day, including determining the protein concentration of the fractions after they have been re-buffered. This is what we got:
- Team Shuttle Vector:
- Genomic DNA isolation of yeast cells with integrated BBa_K863204::GFP construct was done with the Kit. For genotype characterisation an PCR with the praimer pair 5AOX-Genotyp-FW and TT-Genotyp-RV was done and the fragment size was determined.
- Team Fungal and Plant Laccases:
- YPD liquid culture was inoculated with one yeast colony (BBa_K863204::TV5) and incubated at 30°C and 150 rpm.
- Team Cellulose Binding Domain:
- Picked colonies of B0034 and J23100 and plated them on AMP selection agar for plasmid isolation
- Team Site Directed Mutagenesis:
- SDM-PCR of the Shuttle vector
- Added DpnI over night
Thursday October 18th
- Team Activity Tests:
- Finally the day had come to measure the activity of our own produced laccases. This activity assay should give information about the enzyme content in every fraction. To make the measurements comparable we applied the same protein amount in every sample. Again, we used our standard activity assay protocol, but this time we used the Britton-Robinson Buffer at pH 5. Also we applied 0.1 mM ABTS and measured over night.
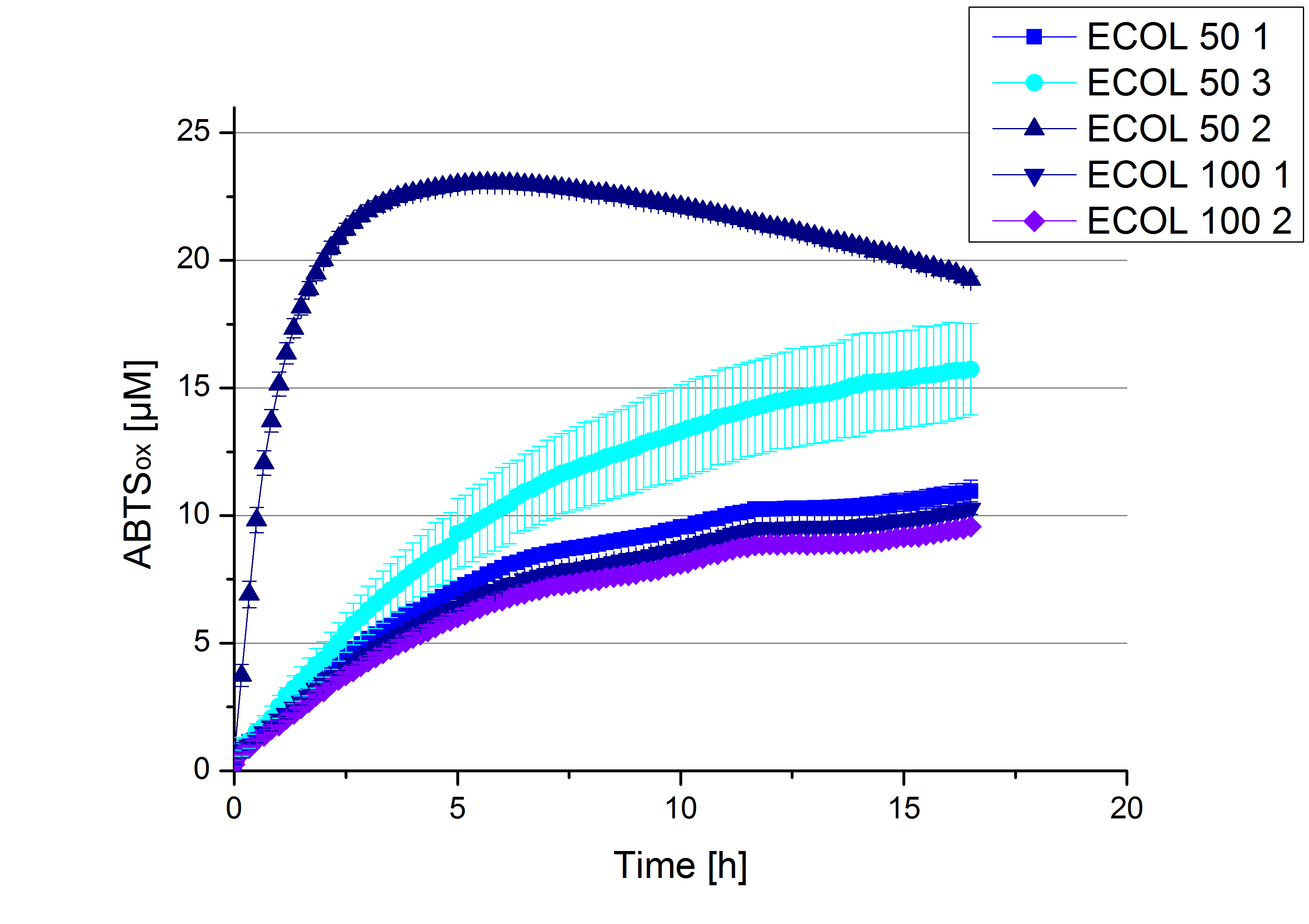
Activity assay of each purified fraction of our new cultivation with ECOL. Samples were re-buffered into H2 and the protein amount in each fraction has been adjusted. The Measurement was done using the standard activity assay protocol over night.
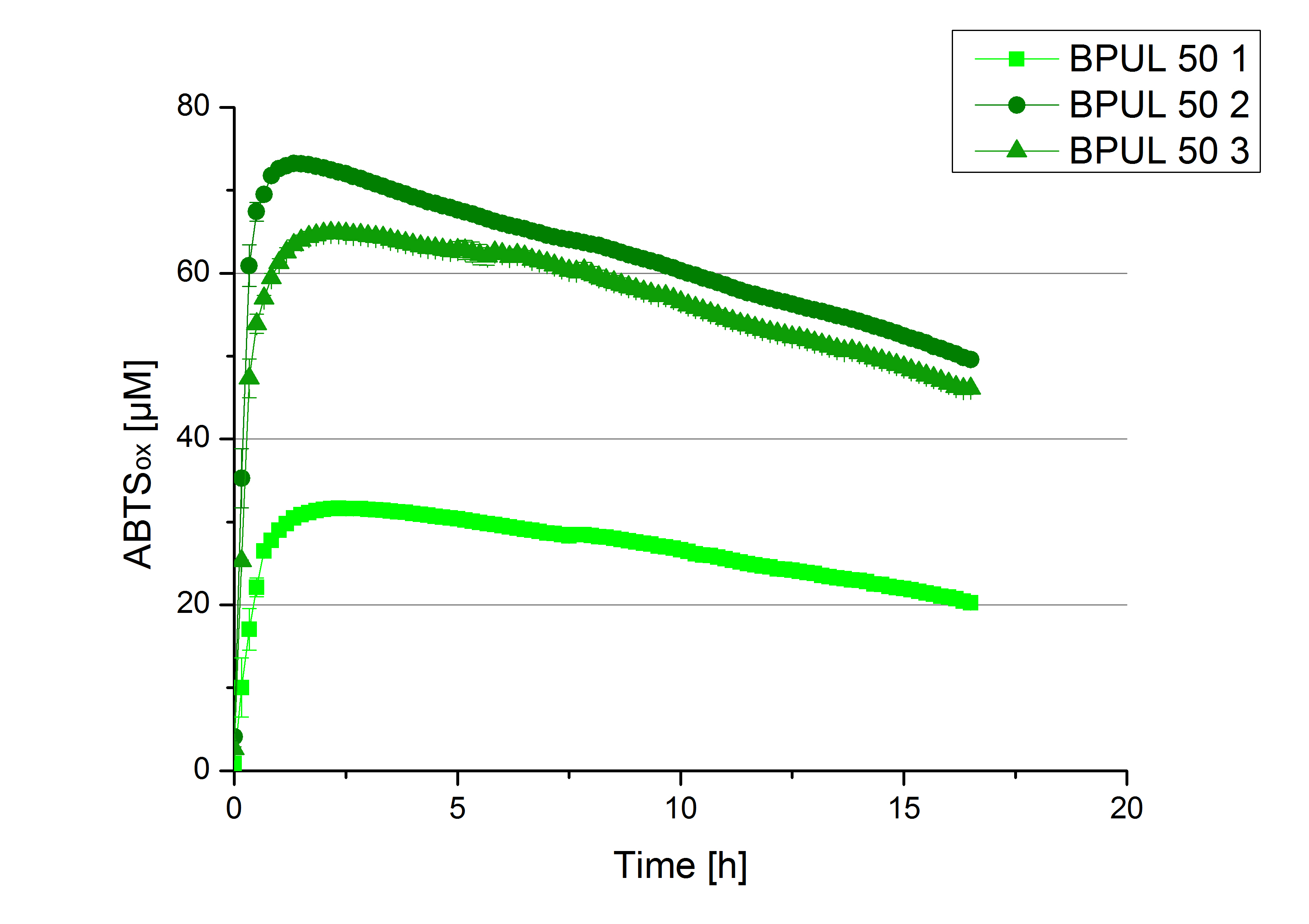
Activity assay of each purified fraction of our new cultivation with BPUL. Samples were re-buffered into H2 and the protein amount in each fraction has been adjusted. The Measurement was done using the standard activity assay protocol over night.
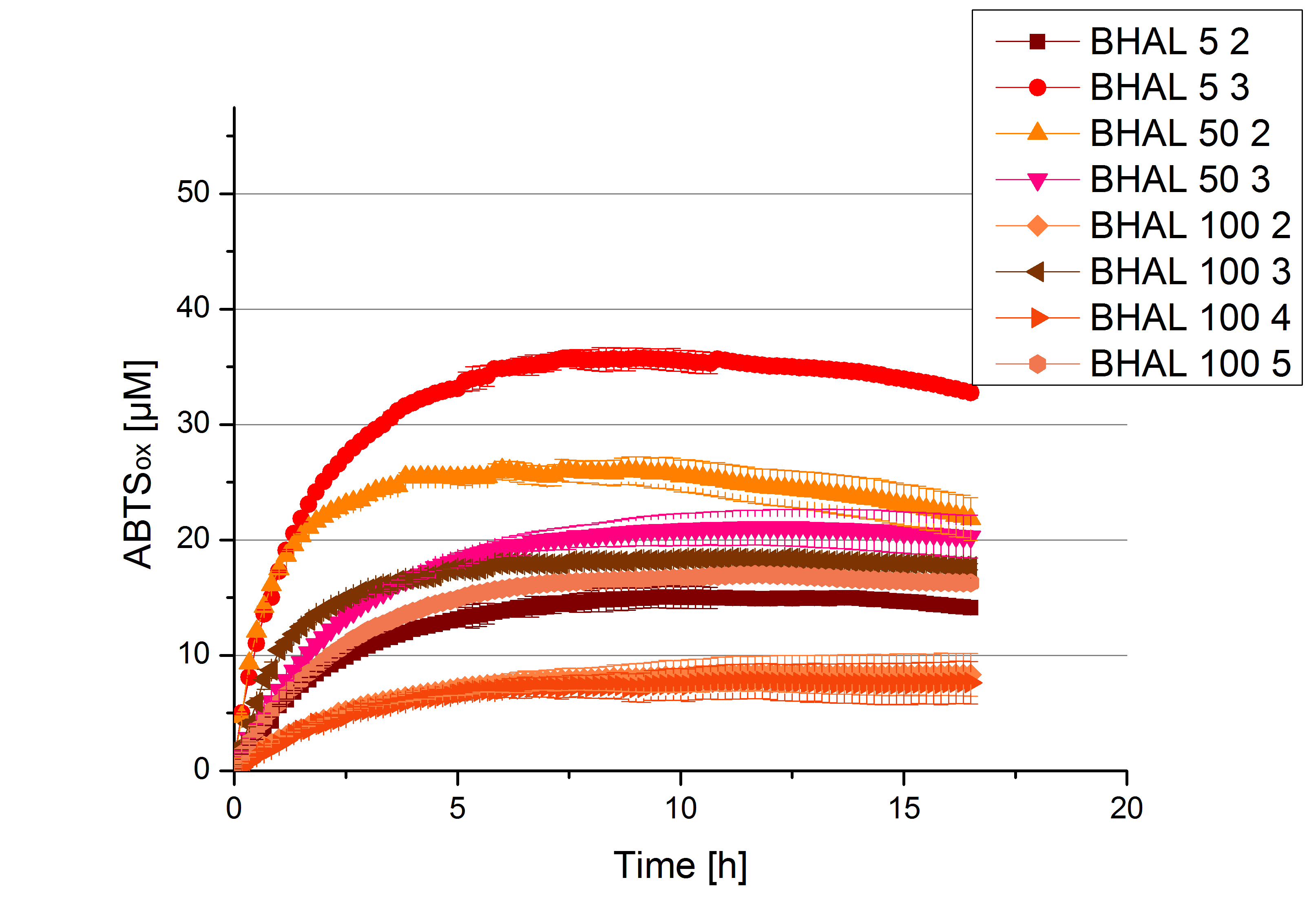
Activity assay of each purified fraction of our new cultivation with BHAL. Samples were re-buffered into H2 and the protein amount in each fraction has been adjusted. The Measurement was done using the standard activity assay protocol over night.
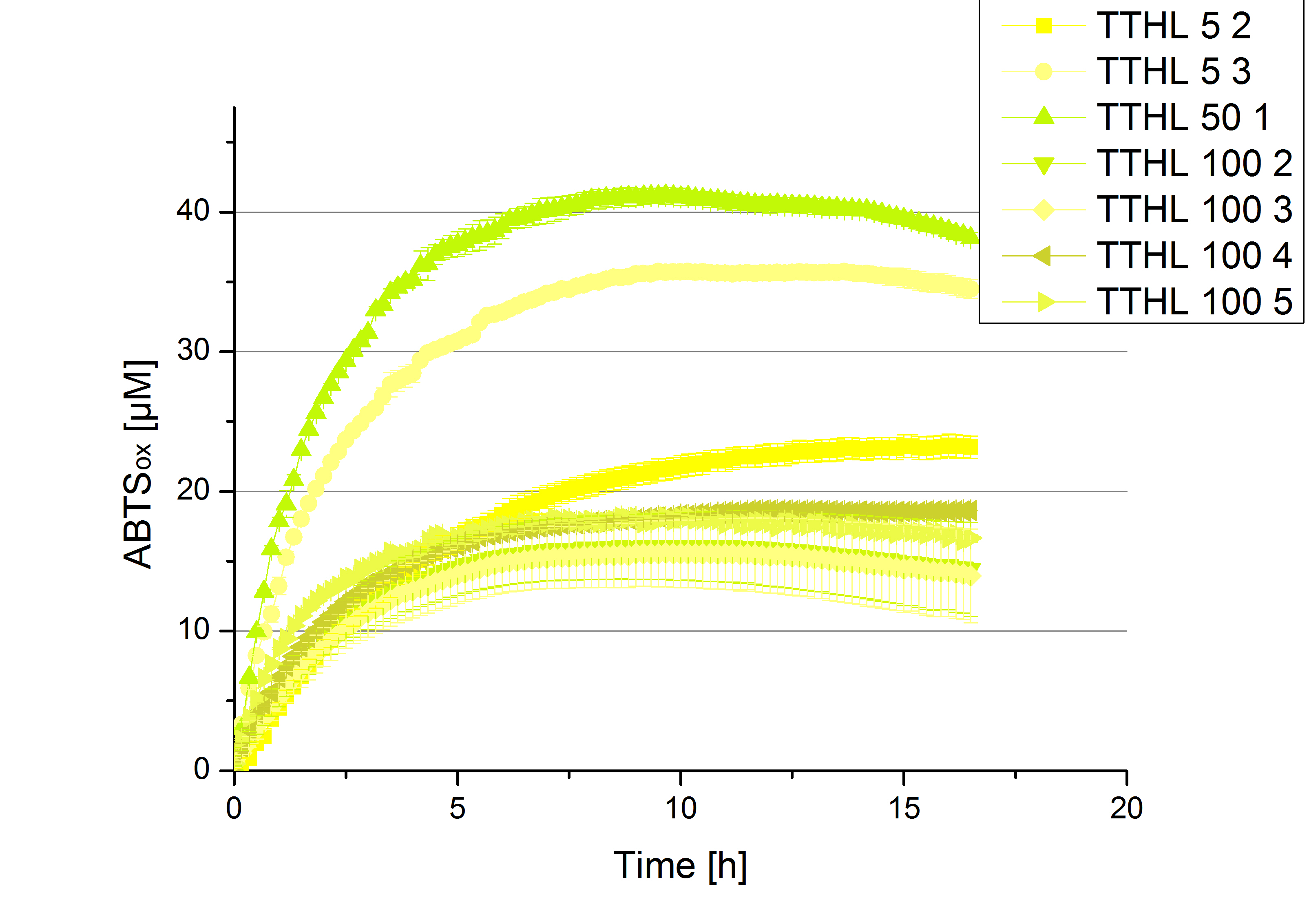
Activity assay of each purified fraction of our new cultivation with TTHL. Samples were re-buffered into H2 and the protein amount in each fraction has been adjusted. The Measurement was done using the standard activity assay protocol over night.
- Team Cellulose Binding Domain:
- Isolated J23100 plasmid
- Digested J23100 with EcoRI and PstI
- Digested pSB1C3-Backbone with EcoRI and PstI
- Clean up of pSB1C3-BB and J23100 + RFP - Insert via Gel
- Ligation of J23100 and pSB1C3-BB
- Isolated B0034 plasmid
- Digested B0034 with SpeI and PstI
- Digested GFP_Freiburg-PCR-product with XbaI and PstI
- Digested an other fraction of the GFP_Freiburg-PCR-product with XbaI and AgeI
- Assembled (Ligation):
- B0034 with GFP_Freiburg
- B0034 with GFP_Freiburg and CBDcex_Freiburg
- B0034 with GFP_Freiburg and CBDclos_Freiburg
- Transformed these three into KRX
- Team Site Directed Mutagenesis:
- Bands of 19C-PRC-product in Gel at 7 kbp (which is correct)
- Clean-Up of 19C-PRC-product
- Transformation of the 19C-PRC-product in XL1 Blue
Friday October 19th
- Team Activity Tests:
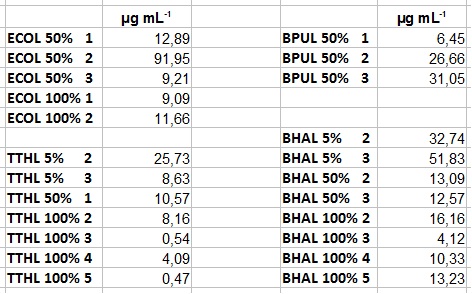
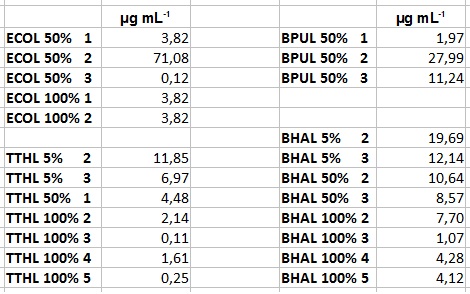
- After getting our interesting activity measurement results, we had to figure out which fraction is going to be used in the following experiments and how much laccase it contains besides other proteins. We agreed, that the most active fraction contains 90 % laccase, which is commonly used in the literature. Since we applied the same protein amount of each fraction for the activity measurements, we made sure that the results really correspond to the amount of laccase. These are the most active fractions we have chosen with their contained laccase amount:
- ECOL: Fraction 50% 2 with a laccase concentration of 63,9 µg mL-1.
- BPUL: Fraction 50% 2 with a laccase concentration of 25,1 µg mL-1.
- BHAL: Fraction 5% 3 with a laccase concentration of 10,9 µg mL-1.
- TTHL: The best fraction of our TTHL laccase was fraction 50% 1 with a laccase concentration of 4,03 µg mL-1. In regard to Team Immobilization and Team Substrate Analysis we had to reach a higher amount of TTHL laccase. So we thought of combining two fractions. We chose the second best fraction of TTHL, which is 5% 3. To calculate the amount of contained laccase we had to compare the activity and reconsidered the slope. Since we stated the fraction with its highest activity as 90 % contained laccase, we calculated the amount of laccase regarding to that one. The following figure shows the slopes we got out of the activity from fraction 50% 1 and fraction 5% 3 (which is the second best). Accordingly to this, fraction 5% 3 contains 68,9% of laccase, which is 4,8 µg mL-1. In total, the composed fraction contains 4,4 µg mL-1.
- After getting our interesting activity measurement results, we had to figure out which fraction is going to be used in the following experiments and how much laccase it contains besides other proteins. We agreed, that the most active fraction contains 90 % laccase, which is commonly used in the literature. Since we applied the same protein amount of each fraction for the activity measurements, we made sure that the results really correspond to the amount of laccase. These are the most active fractions we have chosen with their contained laccase amount:
- Team Cellulose Binding Domain:
- Colony-PCR of B0034 + GFP_Freiburg; B0034 + GFP_Freiburg + CBDcex; B0034 + GFP_Freiburg + CBDcex_Freiburg; B0034 + GFP_Freiburg + CBDclos_Freiburg.
- All negative
- Restriction of Ecol_Freiburg with XbaI, AgeI and DpnI
- Colony-PCR of B0034 + GFP_Freiburg; B0034 + GFP_Freiburg + CBDcex; B0034 + GFP_Freiburg + CBDcex_Freiburg; B0034 + GFP_Freiburg + CBDclos_Freiburg.
- Team Site Directed Mutagenesis:
Saturday October 20th
Sunday October 21st
- Team Cellulose Binding Domain:
- Colonies PCRs of B0034 + GFP_Freiburg; B0034 + GFP_Freiburg + CBDcex_Freiburg; B0034 + GFP_Freiburg + CBDclos_Freiburg
- All negativ
- Transformed once again (with an mix of old an new GFP_Freiburg)
- Isolated plated colonies of transformed colonies (two GFP; two GFP+Cex; two GFP+Clos
- Restriction analysis showed only bands at 2K bp (pSB1C3+BBa_B0034 vector without insert)
- Colonies PCRs of B0034 + GFP_Freiburg; B0034 + GFP_Freiburg + CBDcex_Freiburg; B0034 + GFP_Freiburg + CBDclos_Freiburg
- Team Site Directed Mutagenesis:
- Isolated the plasmids of the six SDM-dishes
- Over-night digestion with NotI
 "
"