Team:Bielefeld-Germany/Results/london
From 2012.igem.org
Summary
Contents |
Introduction
In Munich at the CAS conference we met the iGEM team UCL London for the first time. Discussing about the different projects, a common ground was found. Since the UCL team also produces laccases, but for a different approach, a collaboration has been delevoped. The idea was that each team will characterize the laccase(s) of the other team, to reproduce the results. Therefore the plasmids containing the BioBrick were exchanged. The iGEM UCL London characterized the BioBrick <partinfo>K863005</partinfo> and we characterized <partinfo>K729002</partinfo>. The same laccase CueO from E. coli BL21 (DE3), but different expression vectors and chassis were used by both teams. Therefore the established methods for production and cell disruption had to be discussed and adjusted. For the characterization of the activity, each team used its own method.
For the characterization, the BioBrick <partinfo>K729002</partinfo> was sequenced. It was produced in E. coli KRX and a growth kinetic determined. Furthermore a SDS-PAGE as well as an activity test was made.
Sequencing analysis
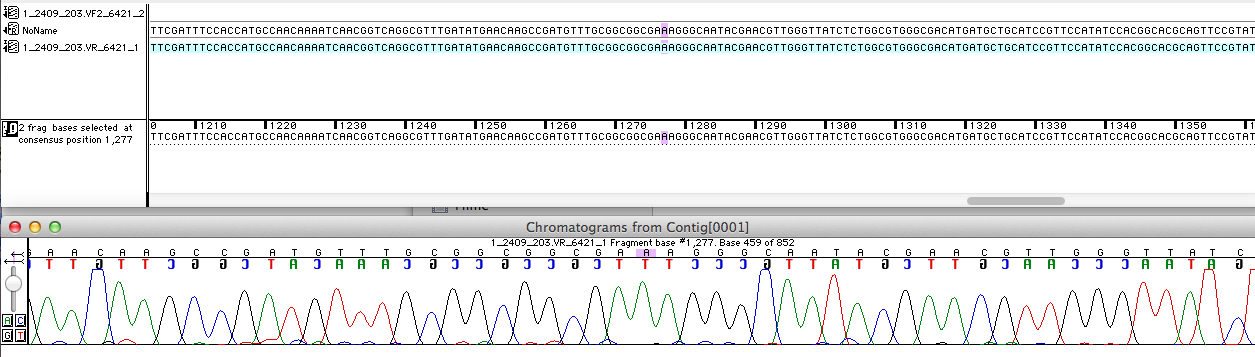
For the improvement of the data collection, the BioBrick of the University College London iGEM team was sequenced. The sequence analysis shows, that the analyzed BioBrick is equal to the sequence published in the partsregistry (see figure 1). Then it was proved if the BioBrick is coding for the E. coli laccase CueO. Therefore the sequence was blasted against CueO from E. coli BL21 (DE3). It matched with 100 %.
Cultivation
E. coli KRX was transformed with the BioBrick <partinfo>K729002</partinfo> of the iGEM team of University College of London. The cells were characterized by the growth kinetics and activity test. E. coli KRX containing <partinfo>K729002</partinfo> was cultivated in LB-medium with a total volume of 60 mL for 12 h at 37 °C (120 rpm) in 250 mL shaking flasks without baffles. To produce a higher amount of the protein, we also cultivated a total volume of 200 mL in a 1 L shaking flask without baffles. The OD600 values were determined every hour. To measure the influence of the transfered BioBrick on the growth of the cells, a negative control (E. coli KRX) was cultivated identically.
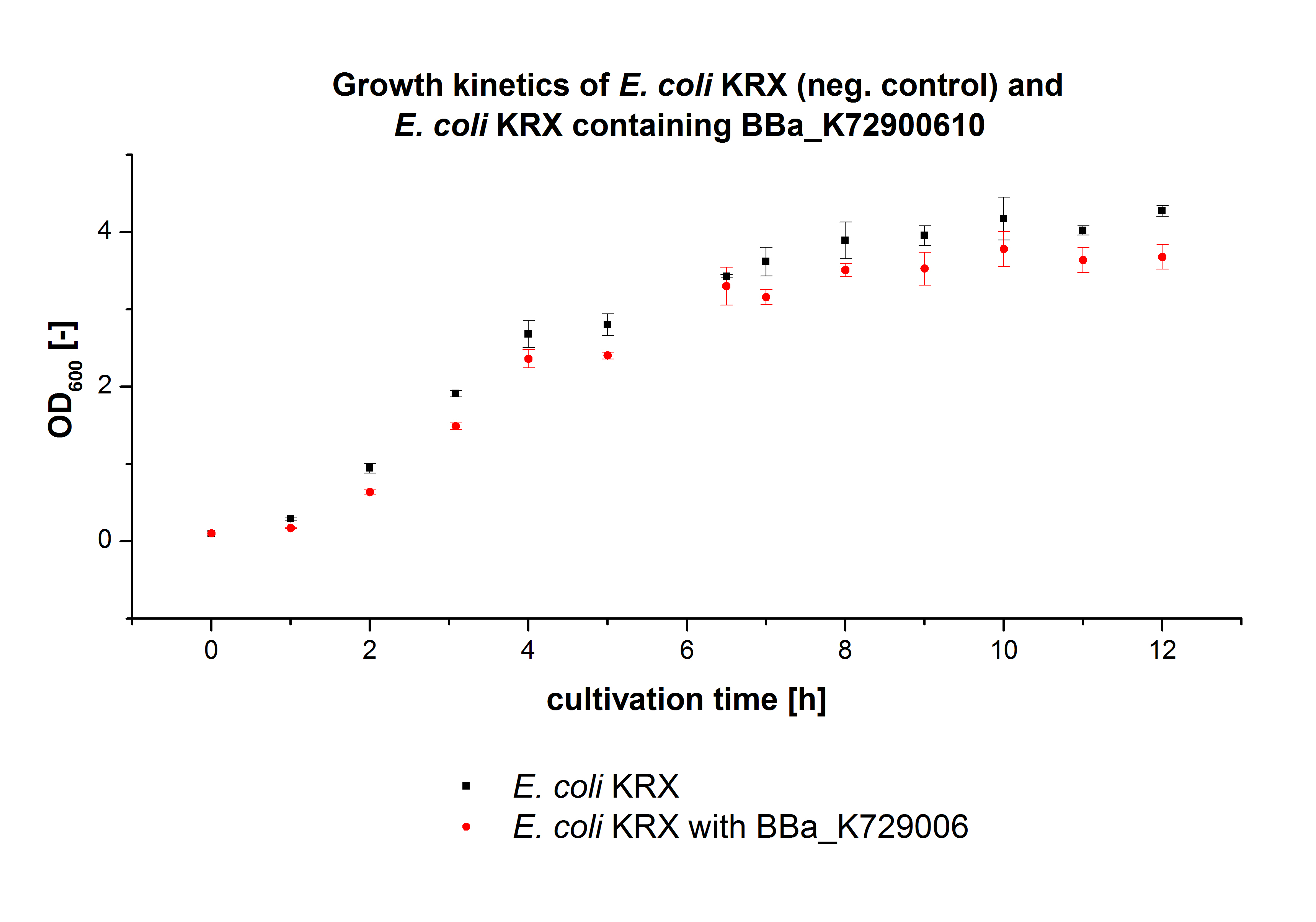
In figure 2 the OD600 of E. coli KRX containing <partinfo>K729002</partinfo> and of the negative control E. coli KRX were plotted. As expected the E. coli KRX containing <partinfo>K729002</partinfo> grew slower than the negative control. This can be explained by the protein expression.
The cells were harvested and centrifuged after 12 h. The pellets were resuspended in 100 mM Na-Acetat-buffer, lysed by sonification and centrifuged. The supernatant was analysed by SDS-PAGE and activity assay.
SDS-PAGE
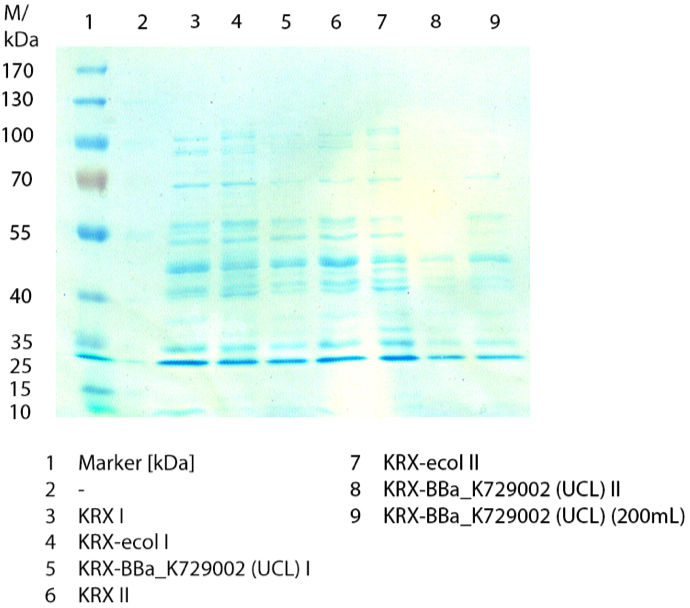
In figure 3 the SDS-PAGE of the lysates of E. coliKRX with <partinfo>BBa_K729006</partinfo>, E. coli KRX (negative control) and furthermore of E. coli KRX with <partinfo>BBa_K863005</partinfo> treated the same way are shown. No clear distinction between the samples is possible. By comparing the bands at 53.4 kDa, the molecular weight of CueO, a small difference of the bandintesity can be suspected. It seems, that the E. coli KRX containig <partinfo>BBa_K863005</partinfo> has a slightly stronger band. The bands at 53.4 kDa were picked and prepared for MALDI-TOF analysis.
Activity Tests
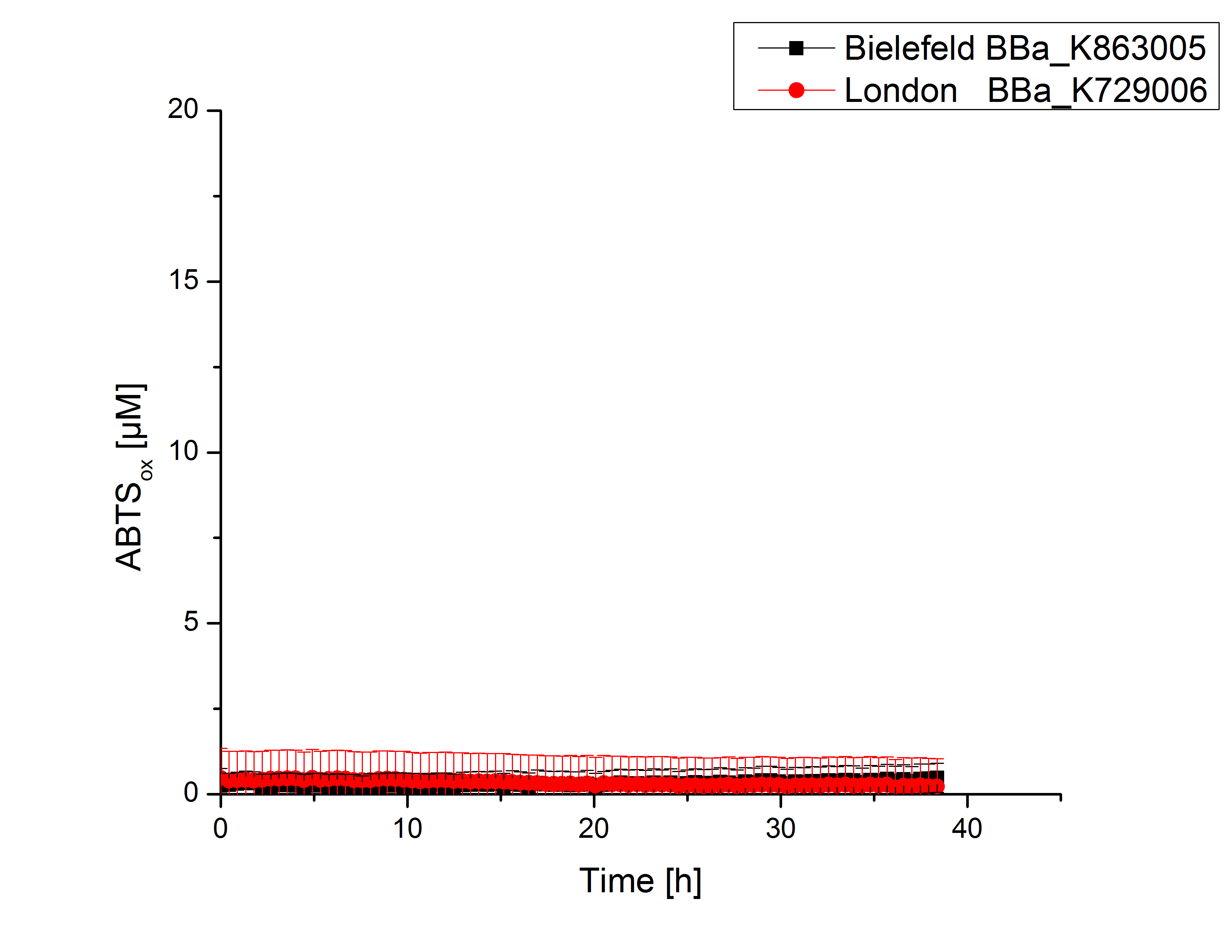
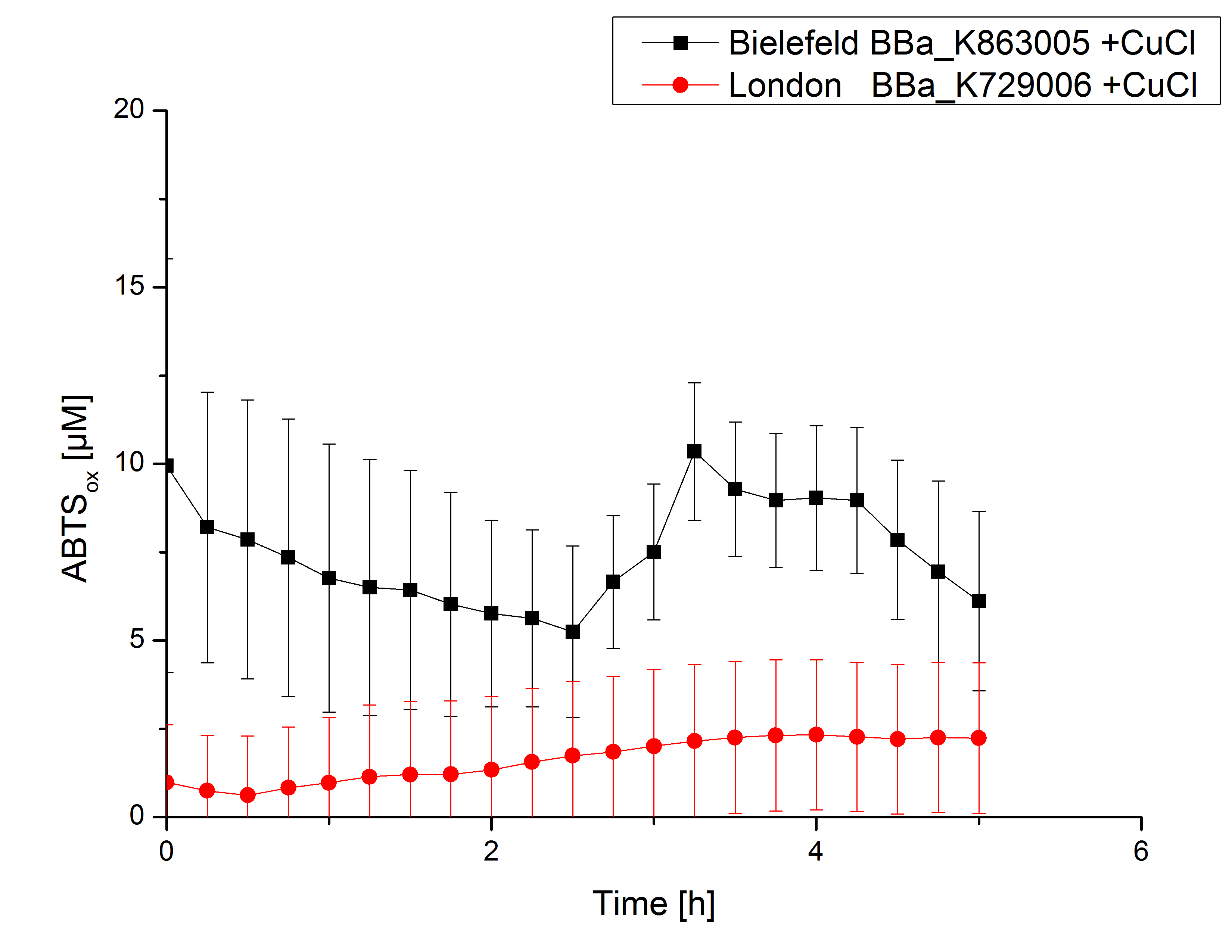
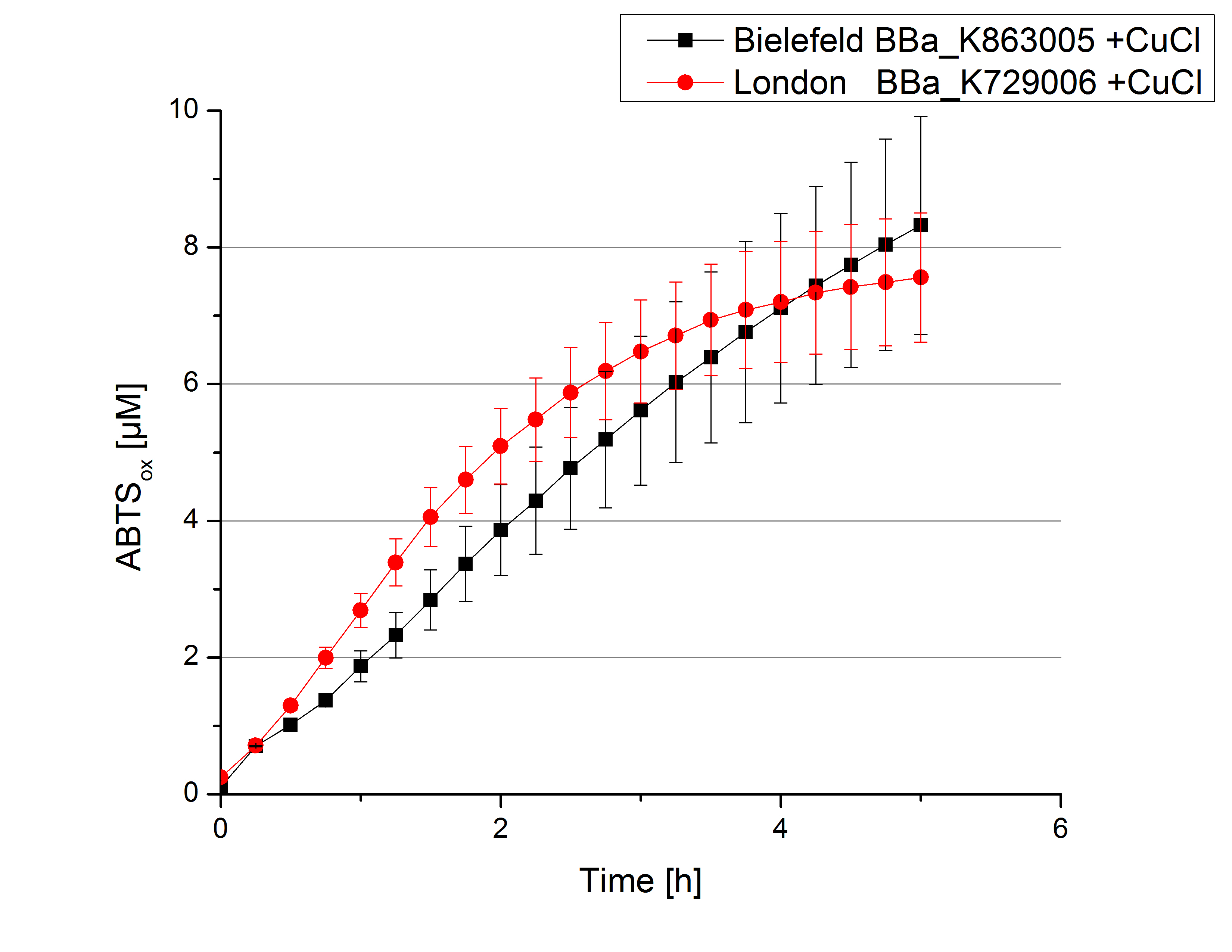
Activity tests were done by using 198 µL of the sample (supernatant of the lysed cells containing <partinfo>K729002</partinfo> and <partinfo>K863005</partinfo>) and 2 µL ABTS (20 mM ABTS stock). Since laccases are capable of oxidizing ABTS the activity of laccases can be detected through measuring the absorption of oxidized ABTS at 420 nm. As positive control a E. coli KRX strain containing <partinfo>K863005</partinfo>, which already has been characterized by the team, was measured too. Using this approach there was no detectable activity over a period of 40 minutes (see figure 4). To gain activity the samples were incubated with 0.4 mM CuCl2 for at least 2 hours. After incubation time another setting was started using 198 µL of the samples containing <partinfo>K729002</partinfo> and <partinfo>K863005</partinfo> and applied 0.1 mM ABTS. The measurements lasted for 5 hours but the results did not show a decrease in oxidized ABTS again (see figure 5). A possible reason can be contaminants in the samples, e.g. salts, low molecular agents, that could affect the activity of the tested laccases. The samples were rebuffered into deionized H2O using HiTrap Desalting Columns and then again incubated with 0.4 mM CuCl2 for 2 hours. This time the measurement setup differed because no buffer remained in the samples. The new measurements were done with 100 mM sodium acetate buffer (pH 5), 158 µL sample and 0.01 mM ABTS for 5 hours at a temperature of 25°C. This preparation of the samples led to oxidizing activity of the laccases and therefore to an increase in oxidized ABTS (see figure 6). After 5 hours both laccases, <partinfo>K729002</partinfo> and <partinfo>K863005</partinfo>, showed great activity and had oxidized ~8 µM ABTS. Since both laccases derive from the same organism they both show a similar reaction behavior.
| 55px | | | | | | | | | | |
 "
"