Team:Bielefeld-Germany/Results/Summary
From 2012.igem.org
| Line 172: | Line 172: | ||
</html> | </html> | ||
zusammenfassung | zusammenfassung | ||
| - | + | <a href="https://2012.igem.org/Team:Bielefeld-Germany/Results/immo">immo</a> | |
| - | + | ||
<html> | <html> | ||
| Line 185: | Line 184: | ||
</html> | </html> | ||
ZUsammenfassung | ZUsammenfassung | ||
| - | + | <a href="https://2012.igem.org/Team:Bielefeld-Germany/Results/substrae">substrae</a> | |
<html> | <html> | ||
| Line 200: | Line 199: | ||
<html> | <html> | ||
zusammenfassung | zusammenfassung | ||
| + | <a href="https://2012.igem.org/Team:Bielefeld-Germany/Results/cbc">cbd</a> | ||
<html> | <html> | ||
| Line 211: | Line 211: | ||
</html> | </html> | ||
A shuttle vector for recombination into the yeast <i>P. pastoris</i> could be developed. With this system it is possible to recombine a protein of interest with a N-terminal mating factor alpha 1 for secretion the protein in the media. This protein of interest could be cloned in frame with one restiction-ligate-cloning-step. The selection depends not on an antibiotic resistance like zeocine, but on a complementation of histidine auxotrophy. | A shuttle vector for recombination into the yeast <i>P. pastoris</i> could be developed. With this system it is possible to recombine a protein of interest with a N-terminal mating factor alpha 1 for secretion the protein in the media. This protein of interest could be cloned in frame with one restiction-ligate-cloning-step. The selection depends not on an antibiotic resistance like zeocine, but on a complementation of histidine auxotrophy. | ||
| + | <a href="https://2012.igem.org/Team:Bielefeld-Germany/Results/vector">read more</a> | ||
<html> | <html> | ||
| Line 225: | Line 226: | ||
The BioBrick [http://partsregistry.org/wiki/index.php?title=Part:BBa_K729002 BBa_K729002] from the [https://2012.igem.org/Team:University_College_London University College London] was characterized by us. Therefore ''E. coli'' KRX containing [http://partsregistry.org/wiki/index.php?title=Part:BBa_K729002 BBa_K729002] and ''E. coli'' KRX as negative control were cultivated in shaking flasks and a growth kinetic was determined. The harvested cells were lysed via [https://2012.igem.org/Team:Bielefeld-Germany/Protocols/Production#Sonification sonification] and the supernatant was purified from substances with a low molecular weight. After purification it was analyzed by [https://2012.igem.org/Team:Bielefeld-Germany/Protocols/Analytics#Sodium_dodecyl_sulfate_polyacrylamide_gel_electrophoresis_.28SDS-PAGE.29 SDS-PAGE], [https://2012.igem.org/Team:Bielefeld-Germany/Protocols/Analytics#Activity_measurements activity assay] as well as [https://2012.igem.org/Team:Bielefeld-Germany/Protocols/Analytics#MALDI MALDI-TOF]. | The BioBrick [http://partsregistry.org/wiki/index.php?title=Part:BBa_K729002 BBa_K729002] from the [https://2012.igem.org/Team:University_College_London University College London] was characterized by us. Therefore ''E. coli'' KRX containing [http://partsregistry.org/wiki/index.php?title=Part:BBa_K729002 BBa_K729002] and ''E. coli'' KRX as negative control were cultivated in shaking flasks and a growth kinetic was determined. The harvested cells were lysed via [https://2012.igem.org/Team:Bielefeld-Germany/Protocols/Production#Sonification sonification] and the supernatant was purified from substances with a low molecular weight. After purification it was analyzed by [https://2012.igem.org/Team:Bielefeld-Germany/Protocols/Analytics#Sodium_dodecyl_sulfate_polyacrylamide_gel_electrophoresis_.28SDS-PAGE.29 SDS-PAGE], [https://2012.igem.org/Team:Bielefeld-Germany/Protocols/Analytics#Activity_measurements activity assay] as well as [https://2012.igem.org/Team:Bielefeld-Germany/Protocols/Analytics#MALDI MALDI-TOF]. | ||
For a comparison ''E. coli'' KRX containing [http://partsregistry.org/wiki/index.php?title=Part:BBa_K863005 BBa_K7863005] was cultivated and also analysed by [https://2012.igem.org/Team:Bielefeld-Germany/Protocols/Analytics#Sodium_dodecyl_sulfate_polyacrylamide_gel_electrophoresis_.28SDS-PAGE.29 SDS-PAGE] and [https://2012.igem.org/Team:Bielefeld-Germany/Protocols/Analytics#Activity_measurements activity assay]. [http://partsregistry.org/wiki/index.php?title=Part:BBa_K729002 BBa_K729002] and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K863005 BBa_K7863005] showed a similar behaviour in oxidizing ABTS. | For a comparison ''E. coli'' KRX containing [http://partsregistry.org/wiki/index.php?title=Part:BBa_K863005 BBa_K7863005] was cultivated and also analysed by [https://2012.igem.org/Team:Bielefeld-Germany/Protocols/Analytics#Sodium_dodecyl_sulfate_polyacrylamide_gel_electrophoresis_.28SDS-PAGE.29 SDS-PAGE] and [https://2012.igem.org/Team:Bielefeld-Germany/Protocols/Analytics#Activity_measurements activity assay]. [http://partsregistry.org/wiki/index.php?title=Part:BBa_K729002 BBa_K729002] and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K863005 BBa_K7863005] showed a similar behaviour in oxidizing ABTS. | ||
| + | <a href="https://2012.igem.org/Team:Bielefeld-Germany/Results/london">read more</a> | ||
<html> | <html> | ||
Revision as of 18:38, 26 September 2012
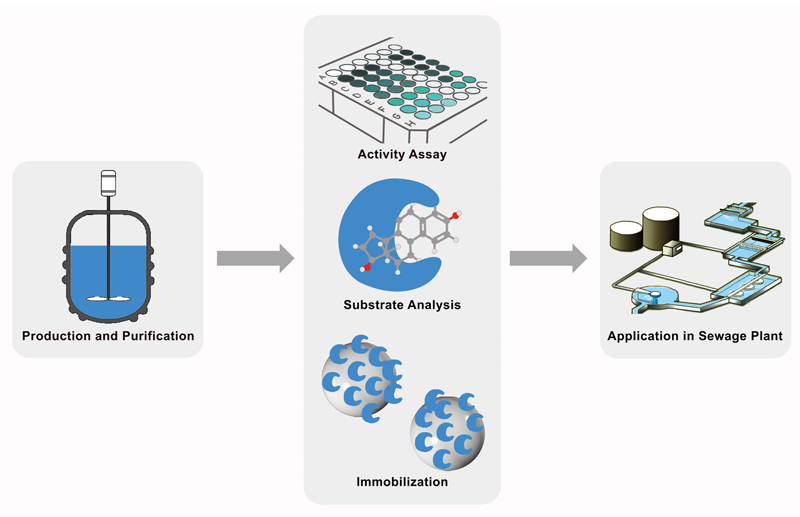
Summary
Cultivation and Purification of the different laccases
During our research we cultivated the following BioBricks and produced several laccase. To simplify the presentation of our results we named the produced laccase like the following system
| Produced and generated BioBricks with the source strain of the DNA-sequence, promoter, protein name and the names given by the iGEM Team Bielefeld | ||||
|---|---|---|---|---|
| BioBrick code | strain | promoter | name of protein | name given by the iGEM Team |
| <partinfo>K863000</partinfo> | Bacillus pumilus DSM 27 | T7 promoter | CotA | BPUL |
| <partinfo>K863005</partinfo> | E. coli BL21(DE3) | T7 promoter | CueO | ECOL |
| <partinfo>K863010</partinfo> | Thermus thermophilus HB27 | T7 promoter | tthL | TTHL |
| <partinfo>K863012</partinfo> | Thermus thermophilus | constitutive promoter (<partinfo>BBa_J23100</partinfo>) | tthL | TTHL |
| <partinfo>K863015</partinfo> | Xanthomonas campestris pv. campestris B100' | T7 | CopA | XCCL |
| <partinfo>K863020</partinfo> | Bacillus halodurans C-125 | T7 | Lbh1 | BHAL |
| <partinfo>K863022</partinfo> | Bacillus halodurans C-125 | constitutive promoter (<partinfo>BBa_J23100</partinfo>) | Lbh1 | BHAL |
All BioBricks of the iGEM Team Bielefeld were screened to identify the best conditions for protein expression. The first trials were made by shaking flask cultivations with different parameters. These parameters were various shaking flask designs , different temperatures, different concentrations of chloramphenicol, various induction strategies , several cultivation times and some cultivations in absence or presence of CuCl2. To detect the produced laccases different analysis methods were performed like SDS-PAGE analysis as well as MALDI-TOF.
Datapage
How our system works
Data for our favorite new parts
- <partinfo>K863000</partinfo> - bpul (laccase from Bacillus pumilus) with T7 promoter, RBS and HIS tag:
- <partinfo>K863005</partinfo> - ecol (laccase from E. coli) with T7 promoter, RBS and HIS tag:
Data for pre-existing parts
We have also characterized the following parts
- <partinfo>BBa_K863012</partinfo> - tthl laccase (from T. thermophilus) with constitutive promoter J23100, RBS and HIS tag:
- <partinfo>BBa_K863022</partinfo> - bhal laccase (from Bacillus halodurans) with constitutive promoter J23100, RBS and HIS tag:
Laccases
Zusammenfassung- <a href="https://2012.igem.org/Team:Bielefeld-Germany/Results/coli">Coli</a>
- <a href="https://2012.igem.org/Team:Bielefeld-Germany/Results/pumi">pumi</a>
- <a href="https://2012.igem.org/Team:Bielefeld-Germany/Results/halo">halo</a>
- <a href="https://2012.igem.org/Team:Bielefeld-Germany/Results/trametis">tramtes</a>
Immobilization
zusammenfassung <a href="https://2012.igem.org/Team:Bielefeld-Germany/Results/immo">immo</a>Subtrate Analytics
ZUsammenfassung <a href="https://2012.igem.org/Team:Bielefeld-Germany/Results/substrae">substrae</a>
Cellulose binding domain
zusammenfassung cbdShuttle vector
A shuttle vector for recombination into the yeast P. pastoris could be developed. With this system it is possible to recombine a protein of interest with a N-terminal mating factor alpha 1 for secretion the protein in the media. This protein of interest could be cloned in frame with one restiction-ligate-cloning-step. The selection depends not on an antibiotic resistance like zeocine, but on a complementation of histidine auxotrophy. <a href="https://2012.igem.org/Team:Bielefeld-Germany/Results/vector">read more</a>
Collaboration with UCL
The BioBrick BBa_K729002 from the University College London was characterized by us. Therefore E. coli KRX containing BBa_K729002 and E. coli KRX as negative control were cultivated in shaking flasks and a growth kinetic was determined. The harvested cells were lysed via sonification and the supernatant was purified from substances with a low molecular weight. After purification it was analyzed by SDS-PAGE, activity assay as well as MALDI-TOF. For a comparison E. coli KRX containing BBa_K7863005 was cultivated and also analysed by SDS-PAGE and activity assay. BBa_K729002 and BBa_K7863005 showed a similar behaviour in oxidizing ABTS. <a href="https://2012.igem.org/Team:Bielefeld-Germany/Results/london">read more</a>
| 55px | | | | | | | | | | |
 "
"