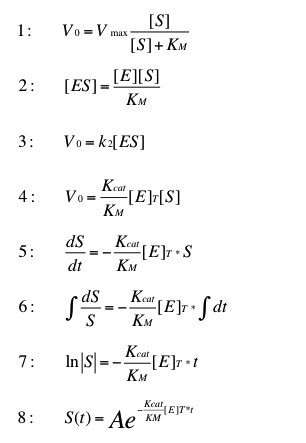Team:Bielefeld-Germany/Modell
From 2012.igem.org
Contents |
Model of a fixed-bed reactor
In our project we plan to construct a fixed-bed reactor, where immobilized laccases degrade synthetic estrogen and other harmful substances. As a small selection we plan to characterize our different laccases for three estrogens, three analgesics, four PAHs, one insecticide and three possible redox mediators. If we could model the degradation of one substrate by one laccase, we could easily replace the specific Kcat KM-1 quotient of other laccases and the amounts of the other substrates.
We ignore the possible cofactors ABTS, syringaldazine and viuloric acid, because on the one hand using these cofactors would increase the costs, and on the other hand they are harmful substances. As shown by Team Substrate Analytic TVEL0 degrades ethinyl estradiol without a redox mediator. So the reaction should follow the Michaelis Menten kinetics.
The reaction
The normal Michaelis Menten kinetic (1) can be adapted for low substrate concentrations. In order to do this formula (2) replaces [ES] in formula (3). The result is formula (4). The velocity of this reaction describes the cange of the substrate concentration over time (5). Formula (6) and (7) solve the integral. Equation (8) is the resulting formula to determine the time dependent substrate concentration by a given startconcentration A.
This transformation of the Michaelis Menten kinetic can be found in "Stryer biochemie". The resulting formula is suitable for very low substrate concentrations. In this case we can estimate degradation of initial substrate concentrations below 0.1 µg L-1.
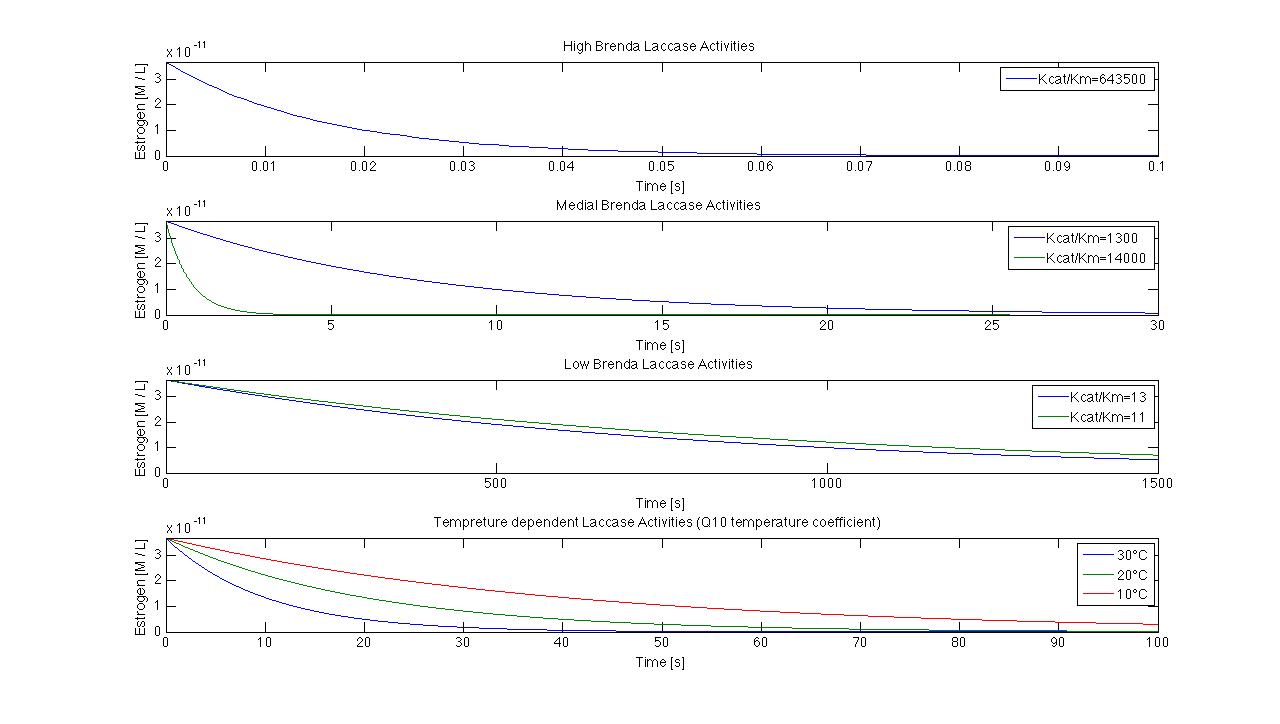
Because we can not find any information about the Kcat KM-1 quotient for the degradation of estradiol in the enzyme database Brenda we decided to use the Kcat KM-1 quotients for the degradation of ABTS, a redox mediator of the laccase. We use ABTS in this case only as placehoder, until we get suitable data for the substrates. In the following picture we show a few possible reactions of our laccase.
We use Kcat KM-1 values from 11 to 643500 that result in degradation time points from 0.07s to 1500s. Furthermore we try to integrate different temperatures to our model. In a span from 10°C to 30°C and a Kcat KM-1 value of 1, the degradation differs from 40s to 100s.
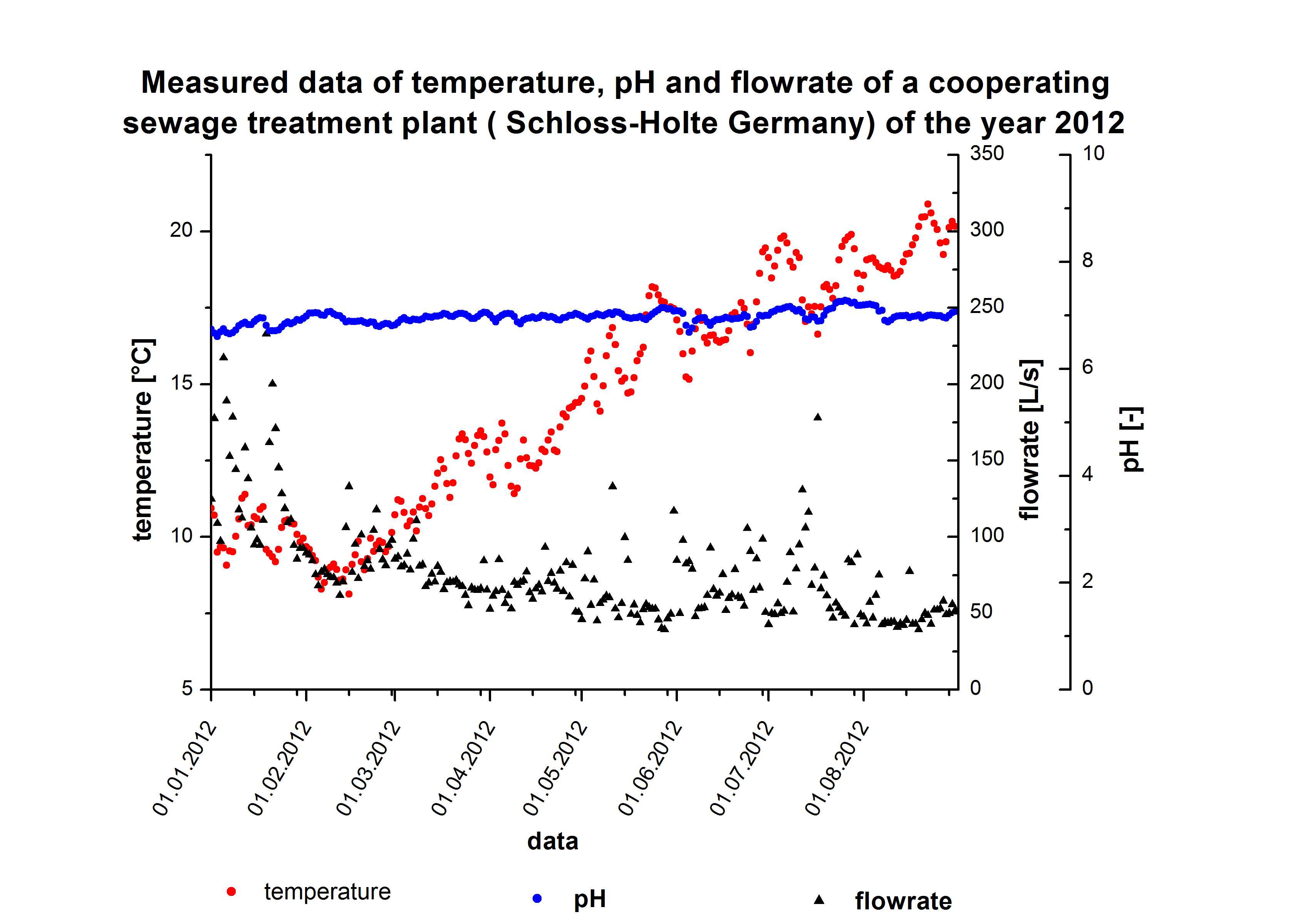
Extend the model with sewageplant data
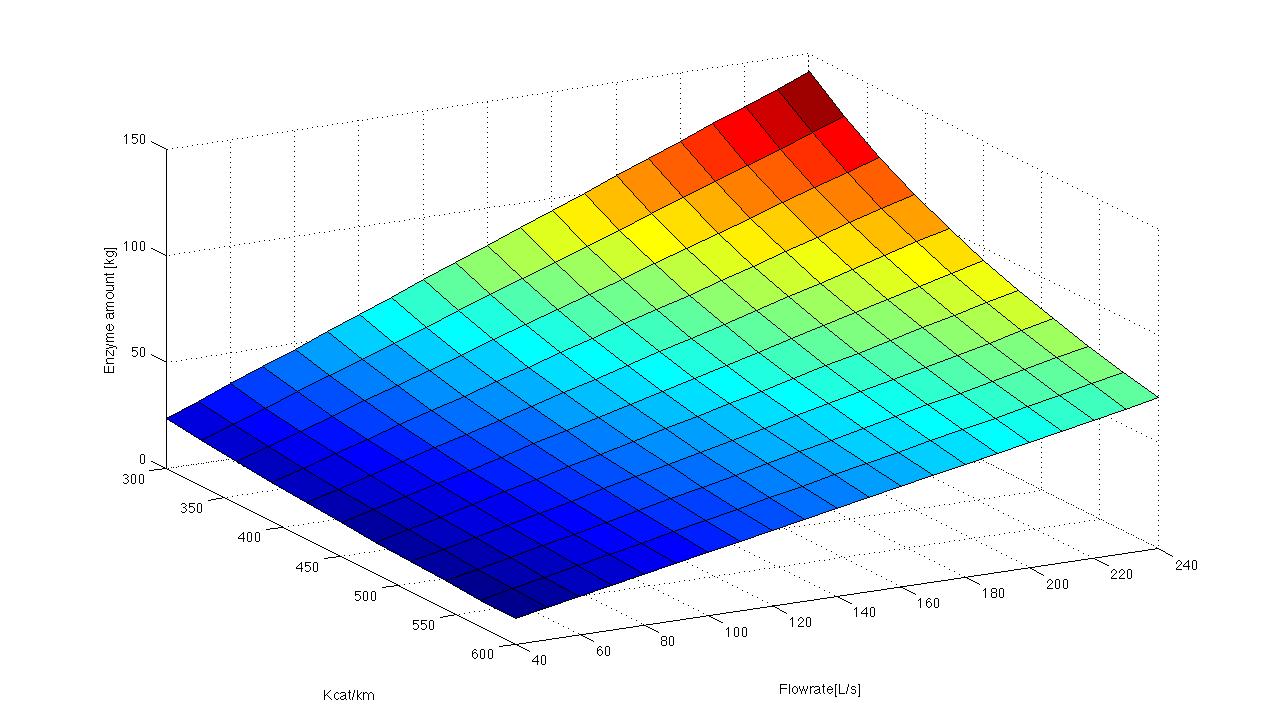
The sewageplant in Schloss Holte provided information about the discharge water, particularly water temperature, pH value and flowrate. The temperature and pH value have a direct influence on the enzymatic activity. Dependent on the enzymatic activity we want to calculate the time our fixed-bed reactor will need to degrdate 80 % of the substrates. This will be the residence time. Combined with the actual flowrate we can determine the reactorsize. To estimate the feasibility we want to know how much enzyme has to be produced for a sewage plant. A 3D model of the required enzme amount dependent of Kcat KM-1 values and flowrate.
Outlook
The next step will be, to add more substrates and different laccases to our model. Hence we will need more data about laccases. Kcat KM-1 values have to be determined as well as the specifity of the laccases. We have the opportunity to work with a lab-scale water treatment system. So we plan to test our model in defined conditions.
| 55px | | | | | | | | | | |
 "
"