Team:LMU-Munich/Bacillus BioBricks
From 2012.igem.org
| Line 143: | Line 143: | ||
<p align="justify">We also tested the [http://partsregistry.org/Promoters/Catalog/Anderson Anderson promoter collection] in ''Bacillus subtilis'' with pSB<sub>Bs</sub>3C-<i>luxABCDE</i>. The data will be online in our [https://2012.igem.org/Team:LMU-Munich/Data Data] section soon.</p> | <p align="justify">We also tested the [http://partsregistry.org/Promoters/Catalog/Anderson Anderson promoter collection] in ''Bacillus subtilis'' with pSB<sub>Bs</sub>3C-<i>luxABCDE</i>. The data will be online in our [https://2012.igem.org/Team:LMU-Munich/Data Data] section soon.</p> | ||
| - | <p align="justify">Furthermore we plan to submit reporter genes optimized for ''Bacillus subtilis'', namely GFP, lacZ, luc and | + | <p align="justify">Furthermore we plan to submit reporter genes optimized for ''Bacillus subtilis'', namely ''GFP'', ''lacZ'', ''luc'' and ''mKate2''. </p> |
Useful for the registry are also 5 tags in Freiburg Standard (cMyc, 10His, Flag, Strep, HA). | Useful for the registry are also 5 tags in Freiburg Standard (cMyc, 10His, Flag, Strep, HA). | ||
| Line 151: | Line 151: | ||
==''Bacillus'' Promoters== | ==''Bacillus'' Promoters== | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
<p align="justify">To get a set of promoters with different strength we characterized several promoters in ''B. subtilis''. They can be divided in three different groups: the constitutive promoters from the [http://partsregistry.org/Part:BBa_J23100 Anderson collection] from the Partsregistry, the constitutive promoters P<sub>''liaG''</sub>, P<sub>''veg''</sub> and P<sub>''lepA''</sub> from ''B. subtilis'', and the inducible promoters P<sub>''liaI''</sub> and P<sub>''xyl''</sub>-''XylR'' from ''B. subtilis''. For the characterization of the different promoters we used the two reporter vectors pSBBs4S-luxABCDE and pSBBs1C-lacZ as well as the reporters lacZ luc and mKate in BioBrick standard.</p> | <p align="justify">To get a set of promoters with different strength we characterized several promoters in ''B. subtilis''. They can be divided in three different groups: the constitutive promoters from the [http://partsregistry.org/Part:BBa_J23100 Anderson collection] from the Partsregistry, the constitutive promoters P<sub>''liaG''</sub>, P<sub>''veg''</sub> and P<sub>''lepA''</sub> from ''B. subtilis'', and the inducible promoters P<sub>''liaI''</sub> and P<sub>''xyl''</sub>-''XylR'' from ''B. subtilis''. For the characterization of the different promoters we used the two reporter vectors pSBBs4S-luxABCDE and pSBBs1C-lacZ as well as the reporters lacZ luc and mKate in BioBrick standard.</p> | ||
| Line 162: | Line 157: | ||
====Anderson promoters==== | ====Anderson promoters==== | ||
| - | <p align="justify">The first group of promoters evaluated are the promoters of the Anderson collection which we call ''Anderson promoters''. They have already been measured in ''Escherichia coli'' where they all showed a constitutive behavior with a different strength. In this project, eleven Anderson promoters were characterized in ''B. subtilis''. Therefore we used the reporter vector pSB<sub>Bs</sub>3C-<i>luxABCDE</i> from the BioBrickBox were they showed quiet a low activity in B. subtilis (see Data). To confirm this result some Anderson promoters were also evaluated in the reporter vector pSB<sub>Bs</sub>1C-''lacZ''(see Data).</p> | + | <p align="justify">The first group of promoters evaluated are the promoters of the [http://partsregistry.org/Part:BBa_J23100 Anderson collection] which we call ''Anderson promoters''. They have already been measured in ''Escherichia coli'' where they all showed a constitutive behavior with a different strength. In this project, eleven Anderson promoters were characterized in ''B. subtilis''. Therefore we used the reporter vector pSB<sub>Bs</sub>3C-<i>luxABCDE</i> from the BioBrickBox were they showed quiet a low activity in B. subtilis (see Data). To confirm this result some Anderson promoters were also evaluated in the reporter vector pSB<sub>Bs</sub>1C-''lacZ''(see Data).</p> |
====Constitutive promoters from ''B. subtilis''==== | ====Constitutive promoters from ''B. subtilis''==== | ||
| - | <p align="justify">The second group of promoters charaterized are constitutive promoters from ''B. subtilis''. We evaluated the promoters P<sub>''liaG''</sub>, P<sub>''veg''</sub> and P<sub>''lepA''</sub>. Therefore we used the reporter vectors pSB<sub>Bs</sub>3C-<i>luxABCDE</i> and pSB<sub>Bs</sub>1C-''lacZ'' as well as the reporter BioBricks lacZ, luc and mKate2.</p> | + | <p align="justify">The second group of promoters charaterized are constitutive promoters from ''B. subtilis''. We evaluated the promoters P<sub>''liaG''</sub>, P<sub>''veg''</sub> and P<sub>''lepA''</sub>. Therefore we used the reporter vectors pSB<sub>Bs</sub>3C-<i>luxABCDE</i> and pSB<sub>Bs</sub>1C-''lacZ'' as well as the reporter BioBricks ''lacZ'', ''luc'' and ''mKate2''.</p> |
| + | |||
| + | *'''P<sub>''liaG''</sub>''' | ||
| + | <p align="justify">PliaG is a weak, constitutive promoter from the LiaRS system, were it is resposible for the production of the components for the resistence against cell wall antibiotics (Reference). This promoter showed a much higher activitx than the Anderson promoters which was still weak in comparison to the other evaluated ''Bacillus'' promoters.</p> | ||
| + | [http://partsregistry.org/wiki/index.php?title=Part:BBa_K823000 BioBrick:BBa_K823000] | ||
| + | |||
| - | |||
Revision as of 09:27, 12 September 2012

The LMU-Munich team is exuberantly happy about the great success at the World Championship Jamboree in Boston. Our project Beadzillus finished 4th and won the prize for the "Best Wiki" (with Slovenia) and "Best New Application Project".
[ more news ]

Bacillus BioBricks
We will create a toolbox of Bacillus BioBricks to contribute to the registry.
This BacillusBioBrickBox contains Bacillus specific:
| Vectors | Promoters | Reporters |
Bacillus Vectors
Here is a list of all the vectors we cloned and used. Please note that pMAD is not in a BioBrick standard, but it is a useful tool to knock out genes in Bacillus subtilis.
For the use of our vectors, please see our Protocols page. A general introduction to Bacillus subtilis and its integrative vectors can be found here. All vectors have ampicillin as E. coli resistance and RFP as selection marker.
| Name BioBrick | E. coli res. | B. subt. res. | Insertion locus | Description | Derived from | Reference |
|---|---|---|---|---|---|---|
| pSBBs1C-lacZ [http://partsregistry.org/wiki/index.php?title=Part:BBa_K823021 (BBa_K823021) ] | Amp | Cam | amyE | with lacZ reporter gene | pAC6 | [http://www.ncbi.nlm.nih.gov/pubmed/11902727 Stülke et al.] |
| pSBBs4S [http://partsregistry.org/wiki/index.php?title=Part:BBa_K823022 (BBa_K823022)] | Amp | Spec | thrC | empty | pDG1731 | [http://www.ncbi.nlm.nih.gov/pubmed/8973347 Guérout-Fleury et al.] |
| pSBBs1C [http://partsregistry.org/wiki/index.php?title=Part:BBa_K823023 (BBa_K823023)] | Amp | Cam | amyE | empty | pDG1662 | [http://www.ncbi.nlm.nih.gov/pubmed/8973347 Guérout-Fleury et al.] |
| pSBBs4S-PXyl [http://partsregistry.org/wiki/index.php?title=Part:BBa_K823024 BBa_K823024] | Amp | Spec | thrC | with Xylose-inducible promoter | pXT | [http://www.ncbi.nlm.nih.gov/pubmed/11069659 Derré et al.] |
| pSBBs3C-luxABCDE [http://partsregistry.org/wiki/index.php?title=Part:BBa_K823025 BBa_K823025] | Amp | Cam | sacA | with luxABCDE reporter cassette | pAH328 | [http://www.ncbi.nlm.nih.gov/pubmed/20709900 Schmalisch et al.] |
| pSBBs0K-Pspac [http://partsregistry.org/wiki/index.php?title=Part:BBa_K823026 BBa_K823026] | Amp | Kan | replicative | with IPTG inducible promoter | pDG148 | [http://www.ncbi.nlm.nih.gov/pubmed/11728721 Joseph et al.] |
| pSBBs2E [http://partsregistry.org/wiki/index.php?title=Part:BBa_K823027 BBa_K823027] | Amp | MLS | lacA | empty | pAX01 | [http://www.ncbi.nlm.nih.gov/pubmed/11274134 Härtl et al.] |
The number in the vector's name codes for the insertion locus and the following letter for the Bacillus subtilis resistance gene according to the following table:
| Number | Insertion locus | Letter | Resistance |
|---|---|---|---|
| 0 | replicative | C | Chloramphenicol |
| 1 | amyE (amylase) | E | MLS (Erythromycin + Lincomycin) |
| 2 | lacA (laccase) | K | Kanamycin |
| 3 | sacA (sucrase) | S | Spectinomycin |
| 4 | thrC (threonine C) |
The concentrations of the antibiotics and the insertion tests can be found in our Protocol section.
The promoters we submit are:
Pveg, PliaI, PliaG, plepA, Pxyl, Pxyl+XylR
We also tested the [http://partsregistry.org/Promoters/Catalog/Anderson Anderson promoter collection] in Bacillus subtilis with pSBBs3C-luxABCDE. The data will be online in our Data section soon.
Furthermore we plan to submit reporter genes optimized for Bacillus subtilis, namely GFP, lacZ, luc and mKate2.
Useful for the registry are also 5 tags in Freiburg Standard (cMyc, 10His, Flag, Strep, HA).
More details to come soon.
Bacillus Promoters
To get a set of promoters with different strength we characterized several promoters in B. subtilis. They can be divided in three different groups: the constitutive promoters from the [http://partsregistry.org/Part:BBa_J23100 Anderson collection] from the Partsregistry, the constitutive promoters PliaG, Pveg and PlepA from B. subtilis, and the inducible promoters PliaI and Pxyl-XylR from B. subtilis. For the characterization of the different promoters we used the two reporter vectors pSBBs4S-luxABCDE and pSBBs1C-lacZ as well as the reporters lacZ luc and mKate in BioBrick standard.
Anderson promoters
The first group of promoters evaluated are the promoters of the [http://partsregistry.org/Part:BBa_J23100 Anderson collection] which we call Anderson promoters. They have already been measured in Escherichia coli where they all showed a constitutive behavior with a different strength. In this project, eleven Anderson promoters were characterized in B. subtilis. Therefore we used the reporter vector pSBBs3C-luxABCDE from the BioBrickBox were they showed quiet a low activity in B. subtilis (see Data). To confirm this result some Anderson promoters were also evaluated in the reporter vector pSBBs1C-lacZ(see Data).
Constitutive promoters from B. subtilis
The second group of promoters charaterized are constitutive promoters from B. subtilis. We evaluated the promoters PliaG, Pveg and PlepA. Therefore we used the reporter vectors pSBBs3C-luxABCDE and pSBBs1C-lacZ as well as the reporter BioBricks lacZ, luc and mKate2.
- PliaG
PliaG is a weak, constitutive promoter from the LiaRS system, were it is resposible for the production of the components for the resistence against cell wall antibiotics (Reference). This promoter showed a much higher activitx than the Anderson promoters which was still weak in comparison to the other evaluated Bacillus promoters.
[http://partsregistry.org/wiki/index.php?title=Part:BBa_K823000 BioBrick:BBa_K823000]
Inducible promoters from B. subtilis
The last group of promoters consists of inducible promoters of B. subtilis e.g. PliaI. They are useful to decide when to turn on gene expression because these promoters need an inducer to start transcription. PliaI can be induced by antibiotics which interact with the lipidII cycle, e.g. bacitracin. In this project this promoter is evaluated like the constitutive promoters with the reporter vector pSBBs3C-luxABCDE which contains the lux operon as a reporter and with the vector pSBBs1C-lacZ which contains the lacZ reporter gene. Therefore the promoters will be amplified from the genome of B. subtilis with primers that contain the restriction sites of the BioBrick standard. Then these inducible promoters will be cloned into the empty vector pSB1C3 to send them to the registry as well as the two reporter vectors to evaluate their strength in B. subtilis. To turn the promoter on we have to add an inducer.
Bacillus Reporters
We designed some reporters that are commonly used in B. subtilis or are codon optimized versions of popular reporter genes. All reporters have a modified iGEM Freiburg standard ([http://partsregistry.org/Help:Assembly_standard_25 RCF 25]) pre- and suffix for assembly of in-frame fusion proteins. Our prefix also includes the B. subitlis optimized RBS.
prefix: GAATTCCGCGGCCGCTTCTAGATAAGGAGGAACTACTATGGCCGGC
suffix: ACCGGTTAATACTAGTAGCGGCCGCTGCAGT
- GFP
We took a gfp derivate of the Bacillus subtilis plasmids pGFPamy and added the BioBrick compatiple pre- and suffix of the Freiburg standard (Assembly 25).
[http://partsregistry.org/wiki/index.php?title=Part:BBa_K823039 BioBrick:BBa_K823039]
- mKate
We synthesized this rfp derivate with a codon-optimized version for the use in B. subtilis with pre- and suffix of the Freiburg standard
[http://partsregistry.org/wiki/index.php?title=Part:BBa_K823029 BioBrick:BBa_K823029]
- LacZ
This lacZ gene is derived from the Bacillus reporter vector pAC6. It is constructed in the Freiburg Standard (Assembly 25) for in-frame fusion proteins. It also includes a Shine-Dalgarno Sequence optimized for Bacillus subtilis translation.
[http://partsregistry.org/wiki/index.php?title=Part:BBa_K823019 BioBrick:BBa_K823019]
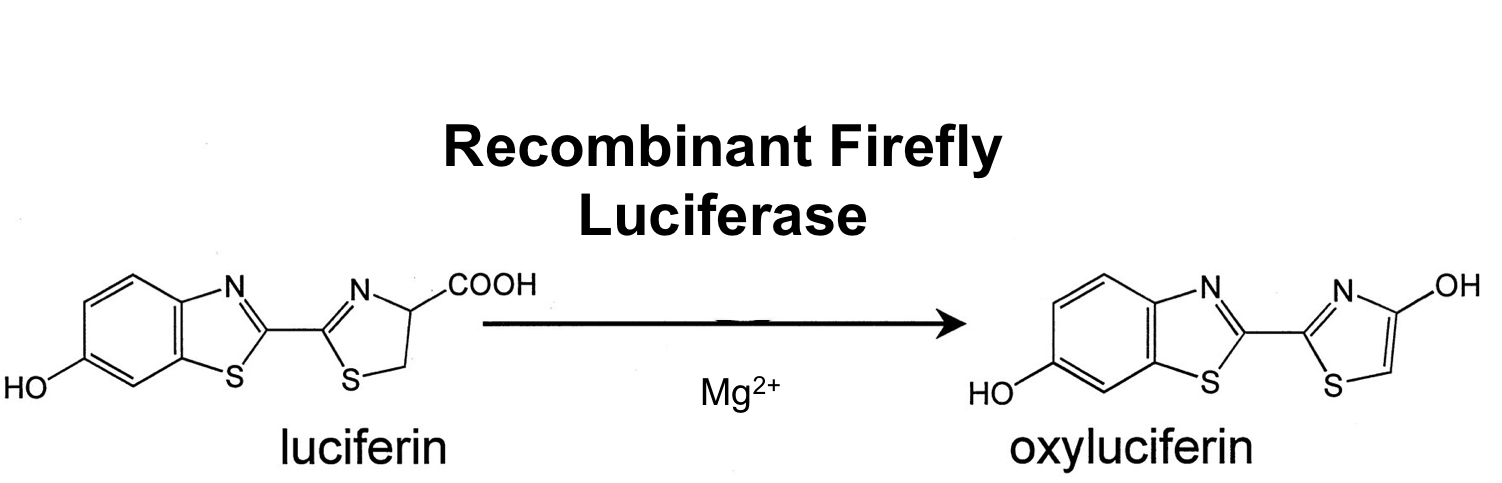
- luc
We synthesized this luciferase for luminescence assays with luciferin as substrate.
[http://partsregistry.org/wiki/index.php?title=Part:BBa_K823028 BioBrick:BBa_K823028]
Project Navigation
| +--+--+--+--+--+--+-- | +--+--+--+--+-- | +--+--+--+--+--+-- | +--+--+--+--+--+-- |

Bacillus BioBrickBOX |

SporeCoat FusionProteins |

Germination STOP |
 "
"