Team:Potsdam Bioware/Project/At a Glance
From 2012.igem.org
(→Antibody Module: Antibody Expression) |
(→Selection Module: Selection or Screening for the Desired Clone) |
||
| (85 intermediate revisions not shown) | |||
| Line 11: | Line 11: | ||
So far, antibodies are typically produced by immunizing mice, sacrificing mice, generating hybridoma by cell fusion and selecting the desired hybridoma clones. This process is very time consuming and the expression quality varies widely. In addition, only natural murine antibodies can be obtained. For modifications or large scale production, the antibodies genes are mostly identified by phage display and then recloned in a defined expression cell line such as CHO. As this is very time consuming, ''E. coli'' based antibody fragment libraries and phage display emerged as a second route. However, in here only scFv or Fab fragments are obtained in the first place, and most applications benefit from the divalent character of the full antibody and need the Fc part. Since the Fc part requires glycosylation for stability a switch from ''E. coli'' to eukaryotic cells is necessary, which again is a laborious step. | So far, antibodies are typically produced by immunizing mice, sacrificing mice, generating hybridoma by cell fusion and selecting the desired hybridoma clones. This process is very time consuming and the expression quality varies widely. In addition, only natural murine antibodies can be obtained. For modifications or large scale production, the antibodies genes are mostly identified by phage display and then recloned in a defined expression cell line such as CHO. As this is very time consuming, ''E. coli'' based antibody fragment libraries and phage display emerged as a second route. However, in here only scFv or Fab fragments are obtained in the first place, and most applications benefit from the divalent character of the full antibody and need the Fc part. Since the Fc part requires glycosylation for stability a switch from ''E. coli'' to eukaryotic cells is necessary, which again is a laborious step. | ||
| - | [[file:UP12_slide2.jpg|920px|center|thumb|''' | + | [[file:UP12_slide2.jpg|920px|center|thumb|'''Figure 1:''' schematic illustration: Overview of the Antibody Generation System]] |
===Our Approach=== | ===Our Approach=== | ||
| Line 33: | Line 33: | ||
==Main Results== | ==Main Results== | ||
| - | ===Antibody Module: Antibody Expression=== | + | ===[https://2012.igem.org/Team:Potsdam_Bioware/Project/Part_Antibody Antibody Module:] Antibody Expression=== |
The aim of the antibody module was to design and assemble antibody constructs that would demonstrate the principle of our generation system. We designed two exemplary antibody constructs: The smaller construct contains a single chain fragment variable domain (scFv) targeting the human EGF-Receptor, a transmembrane domain, a TEV protease cleavage site and an eYFP. The advanced antibody construct consists of a nanobody binding GFP/YFP, a transmembrane domain, a TEV protease cleavage site, mCherry and two LoxP sites. | The aim of the antibody module was to design and assemble antibody constructs that would demonstrate the principle of our generation system. We designed two exemplary antibody constructs: The smaller construct contains a single chain fragment variable domain (scFv) targeting the human EGF-Receptor, a transmembrane domain, a TEV protease cleavage site and an eYFP. The advanced antibody construct consists of a nanobody binding GFP/YFP, a transmembrane domain, a TEV protease cleavage site, mCherry and two LoxP sites. | ||
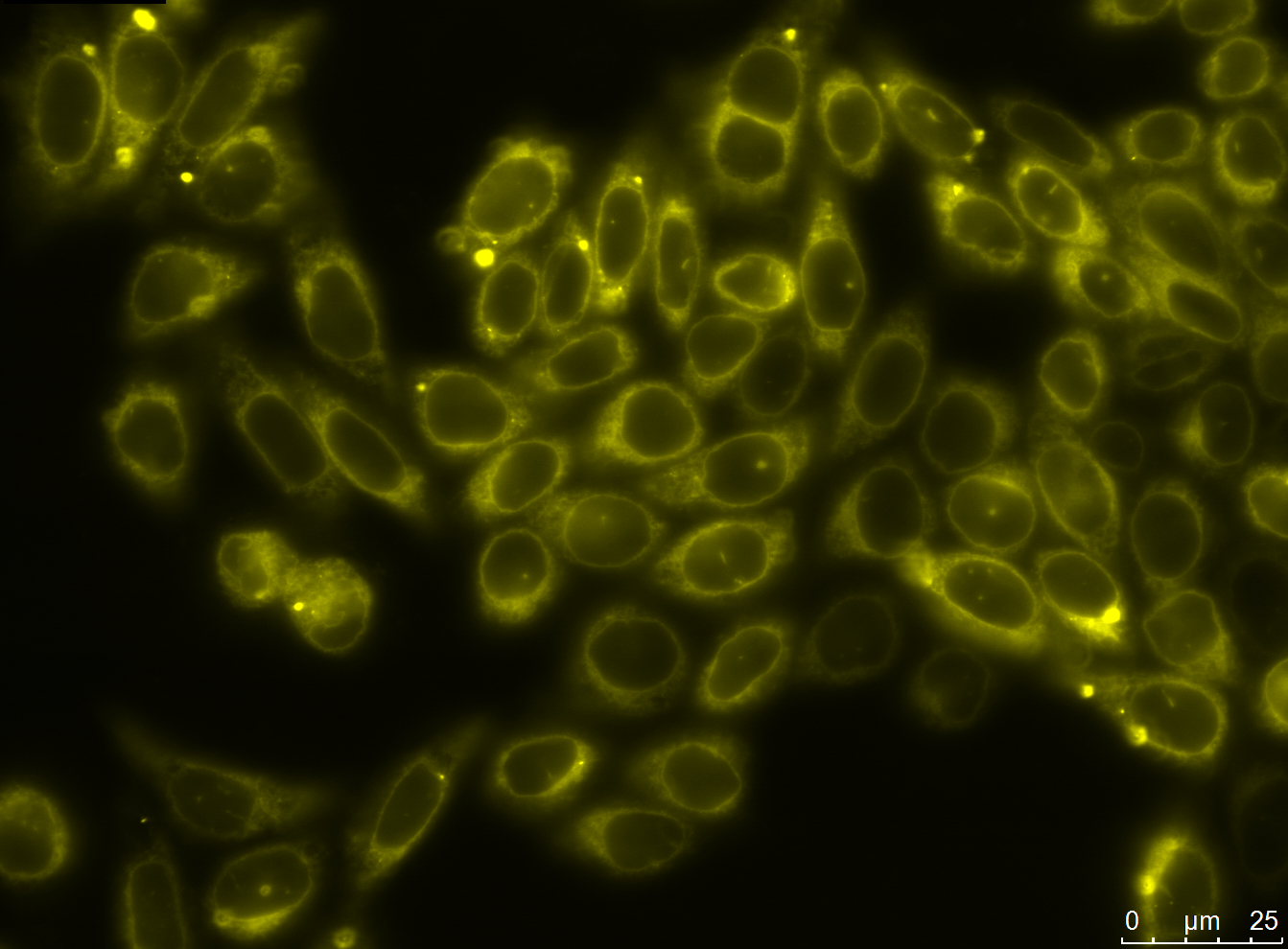
| - | We successfully stably and transiently transfected both antibody constructs in CHO Flp-in cells. With fluorescence microscopy, confocal microscopy, immunfluorescence and FACS we were able to show the expression of the transfected parts. Under certain conditions we have seen a membrane localization of the advanced construct yet we have not been able to generate cells that transport enough molecules to their surface for successful detection. However we have shown that the switch from presenting antibodies to secreting antibodies via the Cre recombinase was successful. | + | We successfully stably and transiently transfected both antibody constructs in CHO Flp-in cells. With fluorescence microscopy, confocal microscopy, immunfluorescence and FACS we were able to show the expression of the transfected parts. Under certain conditions we have seen a membrane localization of the advanced construct yet we have not been able to generate cells that transport enough molecules to their surface for successful detection. However we have shown that the switch from presenting antibodies to secreting antibodies via the Cre recombinase was successful. [https://2012.igem.org/Team:Potsdam_Bioware/Project/Part_Antibody read more] |
<table style="border-spacing:0px"> | <table style="border-spacing:0px"> | ||
| Line 50: | Line 50: | ||
<td>[[file:Up12Haken.png|left|40px|]]</td> | <td>[[file:Up12Haken.png|left|40px|]]</td> | ||
<td>'''membrane localization of advanced construct ([http://partsregistry.org/Part:BBa_K929107 BBa_K929107])'''</td> | <td>'''membrane localization of advanced construct ([http://partsregistry.org/Part:BBa_K929107 BBa_K929107])'''</td> | ||
| - | </tr> | + | </tr><br></table> |
| - | <br> | + | <table style="border-spacing:0px"><tr><td>[[file:UP12_CHO_stabletransfected_Big_Konstruct.jpg|left|395px|thumb|'''Figure 2:''' Fluorescence image of mCherry - advanced antibody construct showing membrane localisation on CHO cells]]</td> |
| - | </table> | + | <td>[[file:UP12_CHO_stabletransfected_clon.jpg|center|460px|thumb|'''Figure 3:''' Fluorescence image of eYFP of the small construct in stably transfected CHO cells]]</table><br> |
| - | + | ||
| - | <table style="border-spacing:0px"> | + | |
| - | <tr> | + | |
| - | <td>[[file:UP12_CHO_stabletransfected_Big_Konstruct.jpg|left| | + | |
| - | <td>[[file:UP12_CHO_stabletransfected_clon.jpg|center| | + | |
| - | + | ||
| - | + | ||
| - | </table> | + | |
| - | <br> | + | |
| - | + | ||
<table style="border-spacing:0px"> | <table style="border-spacing:0px"> | ||
<tr> | <tr> | ||
| Line 69: | Line 59: | ||
</tr> | </tr> | ||
</table> | </table> | ||
| + | <p>The switch from antibody presentation to antibody secretion is induced by the Cre Recombinase. Under the influence of Cre Recombinase the transmembrane domain and mCherry of the advanced antibody construct are eliminated and the antibody is secreted into the supernatant. We demonstrated the functionality of the Cre recombinase via Western Blot. The supernatant was collected and incubated with magnetic beads coupled to GFP via biotin-streptavidin. We detected a band at 41kDa which correlates with the size of the nanobody-Fc construct. [https://2012.igem.org/Team:Potsdam_Bioware/Project/Part_Antibody read more]</p> | ||
| + | [[file:UP12_Cre_Recombinase_Blot_ohneBildunterschrift.png|center|1000px|thumb|'''Figure 4:''' Western Blot of purified Nanobody-Fc from supernatant of cells co-transfected with Cre recombinase and advanced construct. Purification was carried out with magnetic beads. Correct band can be seen at 41kDa]]<br> | ||
| - | + | ===[https://2012.igem.org/Team:Potsdam_Bioware/Project/Part_AID Mutation Module:] Antibody Maturation by Mutation with the AID Enzyme=== | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
'''A'''ctivation '''I'''nduced Cytidine '''D'''eaminase (AID) is one of the key enzymes of antibody maturation in immune system of mammalian organisms. We used the AID to mutate the antibody sequences in CHO cells and in ''E. coli'' during Phage Display. In prior publications, it was shown that the AID is active in CHO cells and ''E. coli'' by using transfection or transformation, respectively.<br> | '''A'''ctivation '''I'''nduced Cytidine '''D'''eaminase (AID) is one of the key enzymes of antibody maturation in immune system of mammalian organisms. We used the AID to mutate the antibody sequences in CHO cells and in ''E. coli'' during Phage Display. In prior publications, it was shown that the AID is active in CHO cells and ''E. coli'' by using transfection or transformation, respectively.<br> | ||
| - | The goals of the mutation module were to express the wildtype and modified AID in CHO cells, to prove the nuclear localization of the modified AID, to determine the mutation rate of wildtype AID and modified AID and to co-transfect the AID with antibody construct. The goal to maturate antibodies was not achieved. | + | The goals of the mutation module were to express the wildtype and modified AID in CHO cells, to prove the nuclear localization of the modified AID, to determine the mutation rate of wildtype AID and modified AID and to co-transfect the AID with antibody construct. The goal to maturate antibodies was not achieved. [https://2012.igem.org/Team:Potsdam_Bioware/Project/Part_AID read more] |
<br> | <br> | ||
| Line 95: | Line 74: | ||
</tr> | </tr> | ||
</table> | </table> | ||
| - | |||
<table> | <table> | ||
<tr> | <tr> | ||
| Line 101: | Line 79: | ||
<td>'''nuclear localisation of modified AID+eGFP [http://partsregistry.org/Part:BBa_K929003 BBa_K929003]'''</td> | <td>'''nuclear localisation of modified AID+eGFP [http://partsregistry.org/Part:BBa_K929003 BBa_K929003]'''</td> | ||
<td></td> | <td></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | <table> | ||
| + | <tr> | ||
<td>[[file:Up12Haken.png|left|40px|]]</td> | <td>[[file:Up12Haken.png|left|40px|]]</td> | ||
<td>'''co-transfection of AID and antibody construct '''</td> | <td>'''co-transfection of AID and antibody construct '''</td> | ||
</tr> | </tr> | ||
</table> | </table> | ||
| - | |||
<table cellpadding="0"> | <table cellpadding="0"> | ||
<tr> | <tr> | ||
<td valign="top"> | <td valign="top"> | ||
| - | [[file:UP_12_intracellulare_location.png|center| | + | [[file:UP_12_intracellulare_location.png|center|505px|thumb|'''Figure 5:''' Overlay image of intracellular localization of modified AID with eGFP]] |
</td> | </td> | ||
<td valign="top"> | <td valign="top"> | ||
| - | [[file:UP_12_cotransfection.jpg|center| | + | [[file:UP_12_cotransfection.jpg|center|350px|thumb|'''Figure 6:''' Overlay image of cotransfection of modified AID with eGFP and antibody-mCherry construct]] |
</td> | </td> | ||
</tr> | </tr> | ||
| Line 122: | Line 103: | ||
<td>'''determination of mutation rates of wildtype and modified AID'''</td> | <td>'''determination of mutation rates of wildtype and modified AID'''</td> | ||
<td></td> | <td></td> | ||
| + | |||
| + | </tr> | ||
| + | </table> | ||
| + | |||
| + | <table> | ||
| + | <tr> | ||
<td>[[file:Up12Haken.png|left|40px|]]</td> | <td>[[file:Up12Haken.png|left|40px|]]</td> | ||
<td>'''recloning of purified antibody plasmids from CHO cells in ''E. coli'' '''</td> | <td>'''recloning of purified antibody plasmids from CHO cells in ''E. coli'' '''</td> | ||
</tr> | </tr> | ||
</table> | </table> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| + | <table><table class = "table center" border=0> | ||
| + | <tr><td><p>To find out the mutation rates of the different AID variant, we tried an unusal setup: recloning of purified antibody plasmids from CHO cells in ''E. coli''. That means that CHO cells were co-transfected with one of the AID constructs (wildtype, modified AID, modified AID+eGFP) and cultured over several days. Afterwards CHO-cells were lysed and plasmid DNA was prepped. To multiply the mutated (and non mutated)plasmids, they were recloned into ''E.coli'' and single clones were sequenced. [https://2012.igem.org/Team:Potsdam_Bioware/Project/Part_AID read more]</p></td><td> | ||
| + | [[file:UP_12_mutation_rate.png|right|400px|thumb|'''Figure 7:''' Comparison of the mutation rates between the wildtype AID, modified AID and modified AID-eGFP in CHO cells (approximately 5000 nucleotides were sequenced)]] | ||
</td> | </td> | ||
</tr> | </tr> | ||
| - | </table> | + | </table><br> |
| - | + | ===[https://2012.igem.org/Team:Potsdam_Bioware/Project/Part_Virus Selection Module:] Selection or Screening for the Desired Clone=== | |
| - | + | ||
| - | + | ||
| - | ===Selection Module: Selection or Screening for the Desired Clone=== | + | |
To select the desired cells from the ones expressing the antibody construct, targeted vital particles are required. Therefore, recombinant adeno-associated virus (rAAV) with yellow fluorescence protein (YFP) on the surface and cyan fluorescence protein (CFP) as Gene of interest was constructed.<br> | To select the desired cells from the ones expressing the antibody construct, targeted vital particles are required. Therefore, recombinant adeno-associated virus (rAAV) with yellow fluorescence protein (YFP) on the surface and cyan fluorescence protein (CFP) as Gene of interest was constructed.<br> | ||
| - | |||
HT1080 were infected with the rAAV. The rAAV with the YFP on the surface and CFP as Gene of interest was used. As the rAAV can infect the HT1080 cells, the HT1080 cells show cyan fluorescence.<br> | HT1080 were infected with the rAAV. The rAAV with the YFP on the surface and CFP as Gene of interest was used. As the rAAV can infect the HT1080 cells, the HT1080 cells show cyan fluorescence.<br> | ||
| - | + | CHO-cells infected with rAAV with the YFP on the surface and CFP as Gene of interest. The CHO-cells can not be infected by the rAAV, which is shown by the yellow fluorescence in the unbleached picture. Bleached CHO-cells dont show YFP fluorescence. [https://2012.igem.org/Team:Potsdam_Bioware/Project/Part_Virus read more]<br> | |
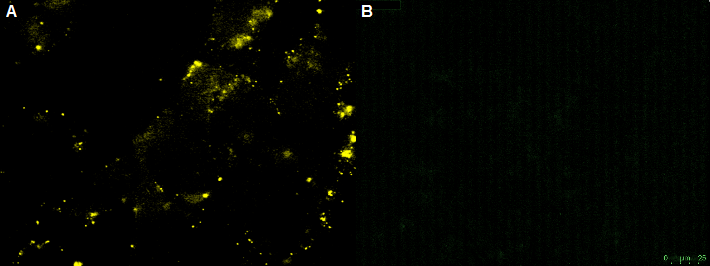
| - | CHO-cells infected with rAAV with the YFP on the surface and CFP as Gene of interest. The CHO-cells can not be infected by the rAAV, which is shown by the yellow fluorescence in the unbleached picture. Bleached CHO-cells dont show YFP fluorescence. | + | |
| - | <br> | + | |
<table style="border-spacing:0px"> | <table style="border-spacing:0px"> | ||
<tr> | <tr> | ||
| Line 154: | Line 132: | ||
</table> | </table> | ||
<br> | <br> | ||
| - | [[File:UP12 HT1080 infected.PNG|left|250px|thumb|''' | + | [[File:UP12 HT1080 infected.PNG|left|250px|thumb|'''Figure 8:''' Fluorescence image of the positive control – infected HT1080 cells with the recombinant adeno-associated virus with YFP on the surface and CFP as the Gene of interest. Cyan fluorescence is the indicator for a successful infection]] |
| - | [[File:UP12 CHO bleaching.PNG|center| | + | [[File:UP12 CHO bleaching.PNG|center|500px|thumb|'''Figure 9:''' Fluorescence image of the negative control - non transfeced CHO-cells with the recombinant adeno-associated virus rAAV). The rAAV are not able to infect the CHO-cells (without nanobody). A) unbleached B) bleached]] |
<br><br><br> | <br><br><br> | ||
| - | |||
<table style="border-spacing:0px"> | <table style="border-spacing:0px"> | ||
<tr> | <tr> | ||
| Line 164: | Line 141: | ||
</tr> | </tr> | ||
</table> | </table> | ||
| + | <center><p>Sortase is an enzyme which catalyzes specific ligation of two proteins to each other. The expressed external ligand-binding 3rd domain of EGFR contains the C-terminal Sortase motif to be recognized by the enzyme Sortase. The coupled protein consisting of 3rd domain of EGFR and the Sortase motif has the size of 24.8 kDa. The N-terminal Sortase motif was fused to the N-terminus of VP2/3 gene of adeno-associated virus to allow the ligation to EGFR by the Sortase. [https://2012.igem.org/Team:Potsdam_Bioware/Project/Part_Virus read more]</p></center>[[File:UP12_mainEGFR.PNG|center|200px|thumb|<b>Figure 10:</b> expression of the 3rd EGFR domain in ''E. coli'' detection was performed using SDS-PAGE and Western-blot]] | ||
| + | <br> | ||
| - | + | ===[https://2012.igem.org/Team:Potsdam_Bioware/Project/Modeling Modeling:] Analyzing and Predicting the Viral Antibody Selection=== | |
| - | + | ||
| - | + | ||
| - | [ | + | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | |||
<table style="border-spacing:0px"> | <table style="border-spacing:0px"> | ||
<tr> | <tr> | ||
| Line 182: | Line 152: | ||
</tr> | </tr> | ||
</table> | </table> | ||
| - | |||
<table> | <table> | ||
<tr><td> | <tr><td> | ||
| + | Modeling is a powerful tool to understand complex interactions or processes. We created a model to illustrate our selection system and to get a better understanding of its behavior under different conditions. Thereby, we wanted to answer two main questions: | ||
| + | |||
| + | #At which time point selection is finished? | ||
| + | #How many antibody presenting cells and how many virus particles do we need at the beginning? | ||
| + | |||
| + | To answer these questions we did both deterministic and stochastic modeling using MATLAB. Furthermore, we did a huge parameter analysis for each model to check what kind of influence each parameter has on our selection system. | ||
| + | |||
| + | Stochastic results showed that the selection is finished on random time points after 100 h or later. Furthermore the analysis of initial concentrations of wt cells and virus revealed that there is an optimum for initial concentrations of wt cells and virus (Figure 11). Both to less and to excessive concentrations prevent the success of the selection system. This optimum is influenced by the other parameters, but if we can determine some of these parameters we can calculate the optimal concentrations to start our selection. [https://2012.igem.org/Team:Potsdam_Bioware/Project/Modeling read more] | ||
| + | |||
| + | </td> | ||
| + | <td> | ||
<table border="1" rules="none"><tr><td> | <table border="1" rules="none"><tr><td> | ||
<html> | <html> | ||
| Line 212: | Line 192: | ||
</html> | </html> | ||
</td></tr><tr><td> | </td></tr><tr><td> | ||
| - | ''' | + | '''Figure 11:''' Dependency of success form initial virus and wt cell concentrations. There are optimal initial concentrations for virus and wt cell |
| - | </td></tr></table> | + | </td></tr></table><br></table><br> |
| - | + | ===[https://2012.igem.org/Team:Potsdam_Bioware/SocialBrick SocialBricks]=== | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
<table style="border-spacing:0px"> | <table style="border-spacing:0px"> | ||
<tr> | <tr> | ||
| Line 241: | Line 205: | ||
<table> | <table> | ||
<tr><td valign="top"> | <tr><td valign="top"> | ||
| - | The idea of SocialBricks is to divide all human practice activities into different parts: the SocialBricks. Here, the SocialBricks stand for every activity which aims to inform people about the Synthetic Biology. We hope that the term SocialBricks will be accepted like the term BioBrick and will be integrated in a registry to show the society what every team has done for more elucidation. | + | The idea of SocialBricks is to divide all human practice activities into different parts: the SocialBricks. Here, the SocialBricks stand for every activity which aims to inform people about the Synthetic Biology. We hope that the term SocialBricks will be accepted like the term BioBrick and will be integrated in a registry to show the society what every team has done for more elucidation. |
| - | For this year we focused on two SocialBricks: “Science meets Politics” and “Science meets People”. For the first one we interviewed politicians from the German parliament: the Bundestag and discussed about Synthetic Biology. For the second part we organized a survey to ask about people's knowledge of Synthetic Biology and for their opinion on this scientific field. We also organized the day of synthetic biology in Potsdam on the main street.<br> | + | For this year we focused on two SocialBricks: “Science meets Politics” and “Science meets People”. For the first one we interviewed politicians from the German parliament: the Bundestag and discussed about Synthetic Biology. For the second part we organized a survey to ask about people's knowledge of Synthetic Biology and for their opinion on this scientific field. We also organized the day of synthetic biology in Potsdam on the main street.<br>The main result of the human practice this year is that the citizens and the politicians see the great potential of Synthetic Biology but also the challenges of this new scientific field. [https://2012.igem.org/Team:Potsdam_Bioware/SocialBrick read more] |
| - | The main result of the human practice this year is that the citizens and the politicians see the great potential of Synthetic Biology but also the challenges of this new scientific field. [https://2012.igem.org/Team:Potsdam_Bioware/SocialBrick | + | |
</td> | </td> | ||
<td> | <td> | ||
| - | [[file:schavan_meeting.jpg| | + | [[file:schavan_meeting.jpg|200px|center|thumb|'''Figure 12:''' Meeting with the Minister of Education and Research: Prof. Dr. Schavan]] |
| - | </td></tr> | + | </td></tr></table><br> |
| - | + | ||
| - | </table> | + | |
<br> | <br> | ||
| - | ===Potsdam Standard | + | ===[https://2012.igem.org/Team:Potsdam_Bioware/Project/Potsdam_Standard Potsdam Standard BBF RFC 91:] a hybrid approach to assemble your gene of interest=== |
<table style="border-spacing:0px"> | <table style="border-spacing:0px"> | ||
<tr> | <tr> | ||
| Line 260: | Line 221: | ||
<tr> | <tr> | ||
<td>[[file:Up12Haken.png|left|40px|]]</td> | <td>[[file:Up12Haken.png|left|40px|]]</td> | ||
| - | <td>'''the | + | <td>'''the correct clones can be seen with naked eye ''' </td> |
</tr> | </tr> | ||
</table> | </table> | ||
The main problems in using the classical approach to assemble different parts with restriction enzymes are on one hand ineffective enzymes with different optimum conditions and on the other hand illegal restriction site in the sequence which restricts the use of standard assemblies.<br> | The main problems in using the classical approach to assemble different parts with restriction enzymes are on one hand ineffective enzymes with different optimum conditions and on the other hand illegal restriction site in the sequence which restricts the use of standard assemblies.<br> | ||
| - | That is the reason why we tried to establish a new RFC with a reduced use of restriction enzymes. This cloning standard is based on the use of thiophosphate primer at the 5’ end for PCR to amplifying the insert. The insert is incubated in iodine/ethanol solution to knock out the 5’ thiophosphates. After that, the new standard cloning vector, developed by us, with a RFP expression cassette as a ligation control is digested with the enzymes Apa I and Sph I. These enzymes generate 3’ overhangs which correspond to the 3’overhang generated by knocking out the thiophosphates. The digested backbone and the pliced insert is mixed, ligated and transformed into E.coli.<br> | + | That is the reason why we tried to establish a new RFC with a reduced use of restriction enzymes. This cloning standard is based on the use of thiophosphate primer at the 5’ end for PCR to amplifying the insert. The insert is incubated in iodine/ethanol solution to knock out the 5’ thiophosphates. After that, the new standard cloning vector, developed by us, with a RFP expression cassette as a ligation control is digested with the enzymes Apa I and Sph I. These enzymes generate 3’ overhangs which correspond to the 3’overhang generated by knocking out the thiophosphates. The digested backbone and the pliced insert is mixed, ligated and transformed into ''E.coli''.<br> |
| - | To proof the new assembly standard, we insert the AID into the new standard cloning vector using the Potsdam Standard. After sequencing, we saw that the cloning was successful without any mutation in the AID sequence. | + | To proof the new assembly standard, we insert the AID into the new standard cloning vector using the Potsdam Standard. After sequencing, we saw that the cloning was successful without any mutation in the AID sequence. [https://2012.igem.org/Team:Potsdam_Bioware/Project/Potsdam_Standard read more]<br> |
| - | [https://2012.igem.org/Team:Potsdam_Bioware/Project/Potsdam_Standard read more] | + | |
<center><table> | <center><table> | ||
<tr> | <tr> | ||
<td> | <td> | ||
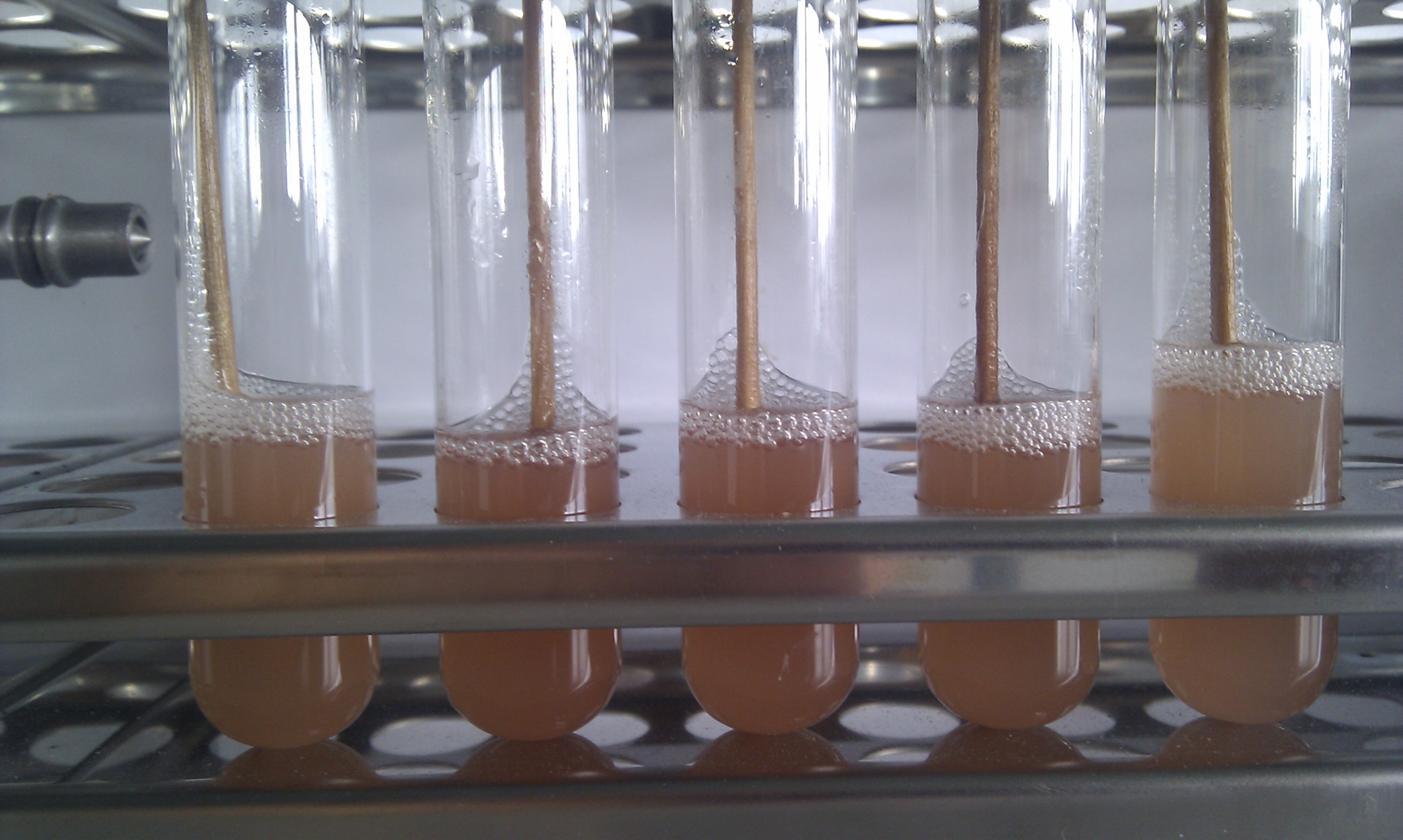
| - | [[file:RFP_culture.jpg|400px|center|thumb|''' | + | [[file:RFP_culture.jpg|400px|center|thumb|'''Figure 13:''' Overnight culture of transformed ''E.coli'' using potsdam standard]] |
<td> | <td> | ||
<td> | <td> | ||
| - | [[File:UP12_RFP_pellets.jpg|400px|center|thumb|''' | + | [[File:UP12_RFP_pellets.jpg|400px|center|thumb|'''Figure 14:''' After centrifugation - red coloured cell pellets]] |
</td> | </td> | ||
</tr> | </tr> | ||
</table> | </table> | ||
Latest revision as of 03:54, 27 October 2012
At a Glance: Antibody Generation System
Antibodies in a Nutshell
The goal of our "Antibody Generation System" is to streamline the generation and production of antibodies by integrating all steps in one cell line. Antibodies are indispensable tools for research and diagnostics and they are used in diverse applications such as ELISA, western blot, affinity purification, and imuno-histology. Antibodies also represent the most important class of biopharmaceuticals with US sales of over 18 billion US$ in 2010 (Aggarwal, 2011) and various therapeutic areas such as cancer, arthritis, and macular degeneration. Depending on the use of the antibodies specific demands must be met. In research, the murine IgG is very typical. For therapy, antibodies need to be 'humanized' to avoid immunological reactions. For some applications downsized or modified versions of antibodies can be used: such as single chain Fv fragments (scFv), Fab fragments, or heavy chain only (camelid) antibodies. However most of the time, the Fc part of the antibody provides crucial functionality like secondary antibody binding in research or cell dependent cytotoxicity in therapy. Most antibodies produced in larger scale are recombinantly expressed in CHO (chinese hamster ovary) cells, due to the fact that CHO cells mimic human glycosylation and can produce large amounts of antibodies. In general, almost 70% of all biopharmaceuticals are produced in CHO cells (CHO consortium, 2012).
State of the Art
So far, antibodies are typically produced by immunizing mice, sacrificing mice, generating hybridoma by cell fusion and selecting the desired hybridoma clones. This process is very time consuming and the expression quality varies widely. In addition, only natural murine antibodies can be obtained. For modifications or large scale production, the antibodies genes are mostly identified by phage display and then recloned in a defined expression cell line such as CHO. As this is very time consuming, E. coli based antibody fragment libraries and phage display emerged as a second route. However, in here only scFv or Fab fragments are obtained in the first place, and most applications benefit from the divalent character of the full antibody and need the Fc part. Since the Fc part requires glycosylation for stability a switch from E. coli to eukaryotic cells is necessary, which again is a laborious step.
Our Approach
We aspire to implement all steps for antibody production the in the well established CHO cells. Our method avoids animal immunization or phage display and the additional recloning and thus bears the potential to be very fast and easy to automatize from start to finish. In addition, we can implement all desired antibody formats large or small and with or without glycosilation. To accomplish our goal, we conceptually divided the steps in modules, which can be tested independently.
The Antibody Module
The design specifications for the antibody module are manifold. The antibody needs to be expressed in CHO cells in a manner that is amenable to selection and maturation as well as soluble/secreted expression. For selection we use cell surface display, which couples phenotype and genotype. Maturation is outlined in the mutations module (see below). And finally for production, we need to implement a genetic switch from cell surface expression to soluble expression. All steps require stable expression over longer periods necessitating stable transfection (i.e. chromosomal integration) of the antibody construct.
We achieved stable integration in CHO cells using the recombination based Flp-In system. As a start we used two different antibody constructs. Our "small antibody construct" gene comprises a signal peptide, a scFv directed against the EGFR (epidermal growth factor receptor), a protease cleavage site, a transmembrane region and as an intracellular reporter a YFP (yellow fluorescent protein). Our advanced antibody construct is a fully synthetic gene comprising a signal peptide, a heavy chain only (nanobody) fragment directed against GFP (green fluorescent protein), an human IgG Fc part, a protease cleavage site a loxP site (which gets translated), a transmembrane region, an mCherry as intracellular reporter and a second loxP site. Both antibody constructs are expressed under the CMV promoter. These two constructs serve to test the system and only at a later stage we plan to generate diversified antibody libraries. The loxP sites in the advanced construct enabled the genetic switch to soluble expression by deleting the transmembrane region and the mCherry reporter upon transfection and expression of the Cre-Recombinase.
The Mutation Module
We expect that stable transfection of antibody libraries is a limiting step in our systems since the up-scale of this process is cumbersome. We estimate that a diversity in the range 106 can be achieved, which is a good starting point to identify lead compounds for binding. However, to generate high affinity antibodies, further maturation might be required. To do so, we utilize the enzyme Activation Induced Cytidine Deaminase (AID), which is one of the key enzymes of antibody maturation in the mammalian immune system. We generated several variants of the AID with deleted nuclear export and added nuclear import sequences and a fused GFP reporter. The resulting mutation rate on the small antibody construct upon co-transfection in CHO-cells was tested to identify the best candidate for mutation. The new constructs localized as expected, and co-expression increased the mutation rate. In comparison, the wt-AID maintained the highest mutation rate. In the next step we will combine maturation with selection.
The Selection Module
Selection is key to the identification of the best binders. Our "selection module" comprises a set of methods to select the desired cell clones from the collection of cells expressing variants of an antibody construct. Depending on the needs, different selection methods might have specific advantages. We set up three methods, whereby two are well established but discontinuous: i) fluorescent activated cell sorting (FACS), which yields many data but requires expensive instrumentation and ii) magnetic beads, which are less expensive and easy to automatize; and last but not least, we develop iii) a novel continuously acting viral selection method. The latter method is based on the assumption that antibody presentation and affinity can be used to modulate viral infection. Virus particles displaying the desired antigen can then either deliver a growth promoting or inhibiting signal. We achieve this by manipulating virus-like particles based on the recombinant Adeno-Associated Virus (rAAV) to present the antigen on the viral surface by either genetic fusion to the capsid protein or by a general enzymatic coupling step using sortase ligation technology. So far we have tested cell sorting (FACS) based on antibody expression, we have coupled YFP, which is the antigen for our test antibody system, to magentic beads, and we have produced rAAV fused at the genetic level to the antigens YFP, CFP and the sortase ligation motif. As a growth inhibiting signal our virus particles deliver a thymidine kinase gene, which enables killing by ganciclovir, and as growth promoting signal a neomycin/geneticin resistance cassette. We are in the process to determine which selection method works the best. As we expect that the viral uptake increases with antibody affinity this system was used in our modeling approaches.
Aggarwal S. What's fueling the biotech engine--2010 to 2011. Nat Biotechnol. 2011 Dec 8;29(12):1083-9. doi: 10.1038/nbt.2060. [http://www.ncbi.nlm.nih.gov/pubmed/22158359 PubMed PMID: 22158359.]
CHO consortium http://hugroup.cems.umn.edu/CHO
Main Results
Antibody Module: Antibody Expression
The aim of the antibody module was to design and assemble antibody constructs that would demonstrate the principle of our generation system. We designed two exemplary antibody constructs: The smaller construct contains a single chain fragment variable domain (scFv) targeting the human EGF-Receptor, a transmembrane domain, a TEV protease cleavage site and an eYFP. The advanced antibody construct consists of a nanobody binding GFP/YFP, a transmembrane domain, a TEV protease cleavage site, mCherry and two LoxP sites.
We successfully stably and transiently transfected both antibody constructs in CHO Flp-in cells. With fluorescence microscopy, confocal microscopy, immunfluorescence and FACS we were able to show the expression of the transfected parts. Under certain conditions we have seen a membrane localization of the advanced construct yet we have not been able to generate cells that transport enough molecules to their surface for successful detection. However we have shown that the switch from presenting antibodies to secreting antibodies via the Cre recombinase was successful. read more
| transiently transfected CHO cell lines with two different antibody constructs (left picture: advanced antibody construct anti-GFP [http://partsregistry.org/Part:BBa_K929107 BBa_K929107]and smaller antibody construct anti-EGFR scFv425 [http://partsregistry.org/Part:BBa_K929101 BBa_K929101] (not shown)) | |
| stably transfected CHO cell lines with two different antibody constructs (advanced antibody construct anti-GFP nanobody [http://partsregistry.org/Part:BBa_K929107 BBa_K929107] (not shown), right picture: smaller antibody construct with anti-EGFR scFv425 [http://partsregistry.org/Part:BBa_K929101 BBa_K929101]) | |
| membrane localization of advanced construct ([http://partsregistry.org/Part:BBa_K929107 BBa_K929107]) |
| proof of functionality of Cre recombinase |
The switch from antibody presentation to antibody secretion is induced by the Cre Recombinase. Under the influence of Cre Recombinase the transmembrane domain and mCherry of the advanced antibody construct are eliminated and the antibody is secreted into the supernatant. We demonstrated the functionality of the Cre recombinase via Western Blot. The supernatant was collected and incubated with magnetic beads coupled to GFP via biotin-streptavidin. We detected a band at 41kDa which correlates with the size of the nanobody-Fc construct. read more
Mutation Module: Antibody Maturation by Mutation with the AID Enzyme
Activation Induced Cytidine Deaminase (AID) is one of the key enzymes of antibody maturation in immune system of mammalian organisms. We used the AID to mutate the antibody sequences in CHO cells and in E. coli during Phage Display. In prior publications, it was shown that the AID is active in CHO cells and E. coli by using transfection or transformation, respectively.
The goals of the mutation module were to express the wildtype and modified AID in CHO cells, to prove the nuclear localization of the modified AID, to determine the mutation rate of wildtype AID and modified AID and to co-transfect the AID with antibody construct. The goal to maturate antibodies was not achieved. read more
| expression of wildtype [http://partsregistry.org/Part:BBa_K929000 BBa_K929000] and modified AID ([http://partsregistry.org/Part:BBa_K929003 BBa_K929003] & [http://partsregistry.org/Part:BBa_K929002 BBa_K929002]) in CHO cells |
| nuclear localisation of modified AID+eGFP [http://partsregistry.org/Part:BBa_K929003 BBa_K929003] |
| co-transfection of AID and antibody construct |
| determination of mutation rates of wildtype and modified AID |
| recloning of purified antibody plasmids from CHO cells in E. coli |
To find out the mutation rates of the different AID variant, we tried an unusal setup: recloning of purified antibody plasmids from CHO cells in E. coli. That means that CHO cells were co-transfected with one of the AID constructs (wildtype, modified AID, modified AID+eGFP) and cultured over several days. Afterwards CHO-cells were lysed and plasmid DNA was prepped. To multiply the mutated (and non mutated)plasmids, they were recloned into E.coli and single clones were sequenced. read more |
Selection Module: Selection or Screening for the Desired Clone
To select the desired cells from the ones expressing the antibody construct, targeted vital particles are required. Therefore, recombinant adeno-associated virus (rAAV) with yellow fluorescence protein (YFP) on the surface and cyan fluorescence protein (CFP) as Gene of interest was constructed.
HT1080 were infected with the rAAV. The rAAV with the YFP on the surface and CFP as Gene of interest was used. As the rAAV can infect the HT1080 cells, the HT1080 cells show cyan fluorescence.
CHO-cells infected with rAAV with the YFP on the surface and CFP as Gene of interest. The CHO-cells can not be infected by the rAAV, which is shown by the yellow fluorescence in the unbleached picture. Bleached CHO-cells dont show YFP fluorescence. read more
| proof of concept: infection by rAAV leads to CFP expression as a selection marker, YFP labeled rAAV is attached to CHO cells |
| expression of sortase motiv+EGFR domain 3 wokrs as a first step to universal virus surface modification |
Sortase is an enzyme which catalyzes specific ligation of two proteins to each other. The expressed external ligand-binding 3rd domain of EGFR contains the C-terminal Sortase motif to be recognized by the enzyme Sortase. The coupled protein consisting of 3rd domain of EGFR and the Sortase motif has the size of 24.8 kDa. The N-terminal Sortase motif was fused to the N-terminus of VP2/3 gene of adeno-associated virus to allow the ligation to EGFR by the Sortase. read more
Modeling: Analyzing and Predicting the Viral Antibody Selection
| The theoretical model supports our assumptions |
|
Modeling is a powerful tool to understand complex interactions or processes. We created a model to illustrate our selection system and to get a better understanding of its behavior under different conditions. Thereby, we wanted to answer two main questions:
To answer these questions we did both deterministic and stochastic modeling using MATLAB. Furthermore, we did a huge parameter analysis for each model to check what kind of influence each parameter has on our selection system. Stochastic results showed that the selection is finished on random time points after 100 h or later. Furthermore the analysis of initial concentrations of wt cells and virus revealed that there is an optimum for initial concentrations of wt cells and virus (Figure 11). Both to less and to excessive concentrations prevent the success of the selection system. This optimum is influenced by the other parameters, but if we can determine some of these parameters we can calculate the optimal concentrations to start our selection. read more |
|
SocialBricks
| Science Meets People & Science Meets Politics |
|
The idea of SocialBricks is to divide all human practice activities into different parts: the SocialBricks. Here, the SocialBricks stand for every activity which aims to inform people about the Synthetic Biology. We hope that the term SocialBricks will be accepted like the term BioBrick and will be integrated in a registry to show the society what every team has done for more elucidation.
For this year we focused on two SocialBricks: “Science meets Politics” and “Science meets People”. For the first one we interviewed politicians from the German parliament: the Bundestag and discussed about Synthetic Biology. For the second part we organized a survey to ask about people's knowledge of Synthetic Biology and for their opinion on this scientific field. We also organized the day of synthetic biology in Potsdam on the main street. |
Potsdam Standard BBF RFC 91: a hybrid approach to assemble your gene of interest
| Potsdam Standard works, new Biobrick was made with Potsdam Standard | |
| the correct clones can be seen with naked eye |
The main problems in using the classical approach to assemble different parts with restriction enzymes are on one hand ineffective enzymes with different optimum conditions and on the other hand illegal restriction site in the sequence which restricts the use of standard assemblies.
That is the reason why we tried to establish a new RFC with a reduced use of restriction enzymes. This cloning standard is based on the use of thiophosphate primer at the 5’ end for PCR to amplifying the insert. The insert is incubated in iodine/ethanol solution to knock out the 5’ thiophosphates. After that, the new standard cloning vector, developed by us, with a RFP expression cassette as a ligation control is digested with the enzymes Apa I and Sph I. These enzymes generate 3’ overhangs which correspond to the 3’overhang generated by knocking out the thiophosphates. The digested backbone and the pliced insert is mixed, ligated and transformed into E.coli.
To proof the new assembly standard, we insert the AID into the new standard cloning vector using the Potsdam Standard. After sequencing, we saw that the cloning was successful without any mutation in the AID sequence. read more
 "
"